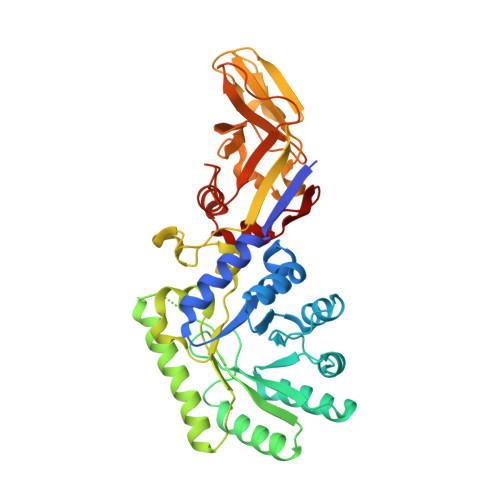
The 1.9 A crystal structure of alanine racemase from Mycobacterium tuberculosis contains a conserved entryway into the active site.
LeMagueres, P., Im, H., Ebalunode, J., Strych, U., Benedik, M.J., Briggs, J.M., Kohn, H., Krause, K.L.(2005) Biochemistry 44: 1471-1481
- PubMed: 15683232 Search on PubMed
- DOI: https://doi.org/10.1021/bi0486583
- Primary Citation Related Structures:
1XFC - PubMed Abstract:
We report the crystal structure of alanine racemase from Mycobacterium tuberculosis (Alr(Mtb)) at 1.9 A resolution. In our structure, Alr(Mtb) is found to be a dimer formed by two crystallographically different monomers, each comprising 384 residues. The domain makeup of each monomer is similar to that of Bacillus and Pseudomonas alanine racemases and includes both an alpha/beta-barrel at the N-terminus and a C-terminus primarily made of beta-strands. The hinge angle between these two domains is unique for Alr(Mtb), but the active site geometry is conserved. In Alr(Mtb), the PLP cofactor is covalently bound to the protein via an internal aldimine bond with Lys42. No guest substrate is noted in its active site, although some residual electron density is observed in the enzyme's active site pocket. Analysis of the active site pocket, in the context of other known alanine racemases, allows us to propose the inclusion of conserved residues found at the entrance to the binding pocket as additional targets in ongoing structure-aided drug design efforts. Also, as observed in other alanine racemase structures, PLP adopts a conformation that significantly distorts the planarity of the extended conjugated system between the PLP ring and the internal aldimine bond.
- Department of Biology and Biochemistry, University of Houston, Houston, Texas 77204-5001, USA.
Organizational Affiliation: