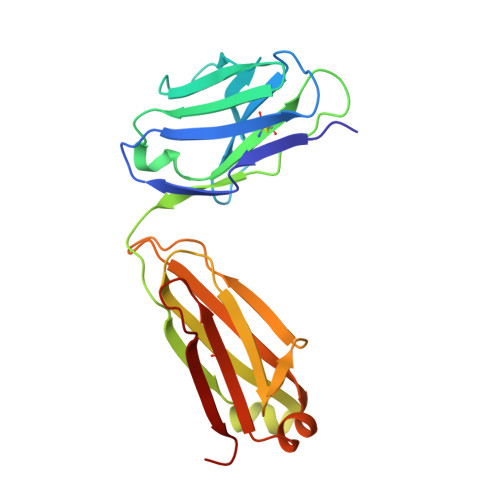
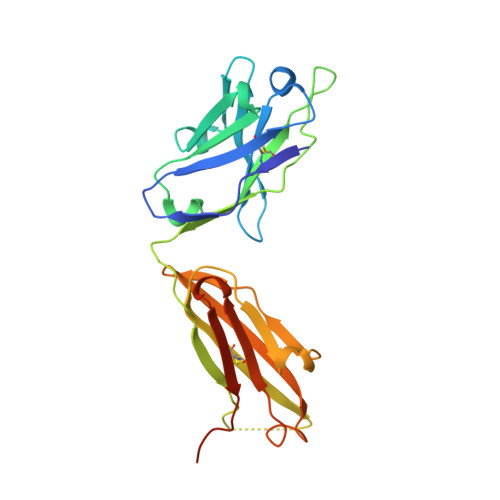
Evidence for Structural Plasticity of Heavy Chain Complementarity-determining Region 3 in Antibody-ssDNA Recognition
Schuermann, J.P., Prewitt, S.P., Davies, C., Deutscher, S.L., Tanner, J.J.(2005) J Mol Biology 347: 965-978
- PubMed: 15784256 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.02.008
- Primary Citation Related Structures:
1XF2, 1XF3, 1XF4 - PubMed Abstract:
Anti-DNA antibodies play important roles in the pathogenesis of autoimmune diseases. They also represent a unique and relatively unexplored class of DNA-binding protein. Here, we present a study of conformational changes induced by DNA binding to an anti-ssDNA Fab known as DNA-1. Three crystal structures are reported: a complex of DNA-1 bound to dT3, and two structures of the ligand-free Fab. One of the ligand-free structures was determined from crystals exhibiting perfect hemihedral twinning, and the details of structure determination are provided. Unexpectedly, five residues (H97-H100A) in the apex of heavy chain complementarity-determining region 3 (HCDR3) are disordered in both ligand-free structures. Ligand binding also caused a 2-4A shift of the backbone of Tyr L92 and ordering of the L92 side-chain. In contrast, these residues are highly ordered in the Fab/dT3 complex, where Tyr H100 and Tyr H100A form intimate stacking interactions with DNA bases, and L92 forms the 5' end of the binding site. The structures suggest that HCDR3 is very flexible and adopts multiple conformations in the ligand-free state. These results are discussed in terms of induced fit and pre-existing equilibrium theories of ligand binding. Our results allow new interpretations of existing thermodynamic and mutagenesis data in terms of conformational entropy and the volume of conformational space accessible to HCDR3 in the ligand-free state. In the context of autoimmune disease, plasticity of the ligand-free antibody could provide a mechanism by which anti-DNA antibodies bind diverse host ligands, and thereby contribute to pathogenicity.
- Department of Chemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: