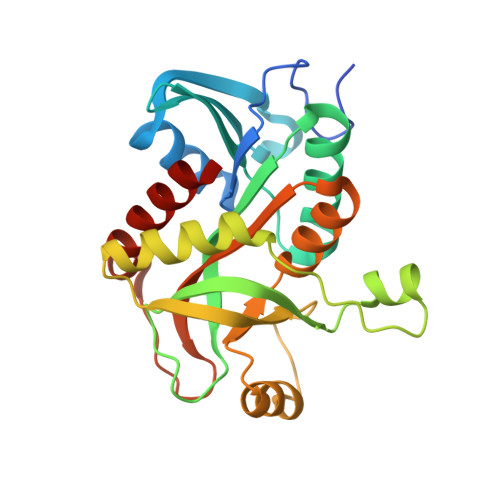
Structure of purine nucleoside phosphorylase (DeoD) from Bacillus anthracis.
Grenha, R., Levdikov, V.M., Fogg, M.J., Blagova, E.V., Brannigan, J.A., Wilkinson, A.J., Wilson, K.S.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 459-462
- PubMed: 16511068 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S174430910501095X
- Primary Citation Related Structures:
1XE3 - PubMed Abstract:
Protein structures from the causative agent of anthrax (Bacillus anthracis) are being determined as part of a structural genomics programme. Amongst initial candidates for crystallographic analysis are enzymes involved in nucleotide biosynthesis, since these are recognized as potential targets in antibacterial therapy. Purine nucleoside phosphorylase is a key enzyme in the purine-salvage pathway. The crystal structure of purine nucleoside phosphorylase (DeoD) from B. anthracis has been solved by molecular replacement at 2.24 A resolution and refined to an R factor of 18.4%. This is the first report of a DeoD structure from a Gram-positive bacterium.
- Department of Chemistry, University of York, York YO10 5YW, England.
Organizational Affiliation: