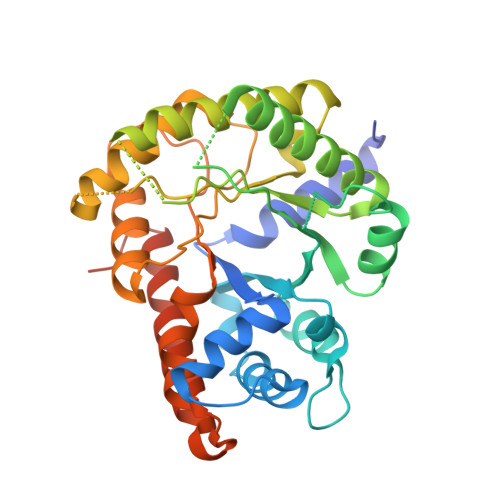
Structure of the thermolabile mutant aldolase B, A149P: molecular basis of hereditary fructose intolerance.
Malay, A.D., Allen, K.N., Tolan, D.R.(2005) J Mol Biology 347: 135-144
- PubMed: 15733923 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.01.008
- Primary Citation Related Structures:
1XDL, 1XDM - PubMed Abstract:
Hereditary fructose intolerance (HFI) is a potentially lethal inborn error in metabolism caused by mutations in the aldolase B gene, which is critical for gluconeogenesis and fructose metabolism. The most common mutation, which accounts for 53% of HFI alleles identified worldwide, results in substitution of Pro for Ala at position 149. Structural and functional investigations of human aldolase B with the A149P substitution (AP-aldolase) have shown that the mutation leads to losses in thermal stability, quaternary structure, and activity. X-ray crystallography is used to reveal the structural basis of these perturbations. Crystals of AP-aldolase are grown at two temperatures (4 degrees C and 18 degrees C), and the structure solved to 3.0 angstroms resolution, using the wild-type structure as the phasing model. The structures reveal that the single residue substitution, A149P, causes molecular disorder around the site of mutation (residues 148-159), which is propagated to three adjacent beta-strand and loop regions (residues 110-129, 189-199, 235-242). Disorder in the 110-129-loop region, which comprises one subunit-subunit interface, provides an explanation for the disrupted quaternary structure and thermal instability. Greater structural perturbation, particularly at a Glu189-Arg148 salt bridge in the active-site architecture, is observed in the structure determined at 18 degrees C, which could explain the temperature-dependent loss in activity. The disorder revealed in these structures is far greater than that predicted by homology modeling and underscores the difficulties in predicting perturbations of protein structure and function by homology modeling alone. The AP-aldolase structure reveals the molecular basis of a hereditary disease and represents one of only a few structures known for mutant proteins at the root of the thousands of other inherited disorders.
- Biology Department, Boston University, Boston, MA 02215, USA.
Organizational Affiliation: