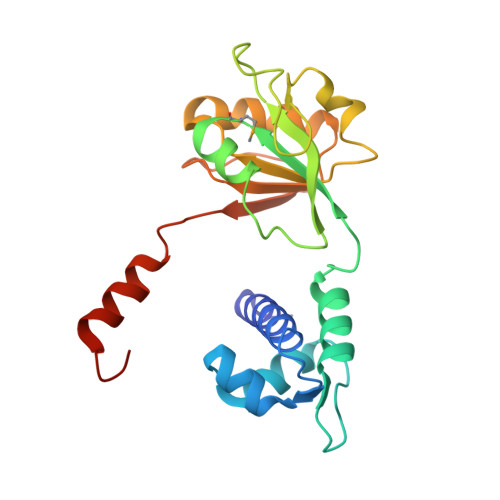
X-Ray Structure of a Rex-Family Repressor/NADH Complex: Insights into the Mechanism of Redox Sensing
Sickmier, E.A., Brekasis, D., Paranawithana, S., Bonanno, J.B., Paget, M.S., Burley, S.K., Kielkopf, C.L.(2005) Structure 13: 43-54
- PubMed: 15642260 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.10.012
- Primary Citation Related Structures:
1XCB - PubMed Abstract:
The redox-sensing repressor Rex regulates transcription of respiratory genes in response to the intra cellular NADH/NAD(+) redox poise. As a step toward elucidating the molecular mechanism of NADH/NAD(+) sensing, the X-ray structure of Thermus aquaticus Rex (T-Rex) bound to effector NADH has been determined at 2.9 A resolution. The fold of the C-terminal domain of T-Rex is characteristic of NAD(H)-dependent enzymes, whereas the N-terminal domain is similar to a winged helix DNA binding motif. T-Rex dimerization is primarily mediated by "domain-swapped" alpha helices. Each NADH molecule binds to the C-terminal domain near the dimer interface. In contrast to NAD(H)-dependent enzymes, the nicotinamide is deeply buried within a hydrophobic pocket that appears to preclude substrate entry. We show that T-Rex binds to the Rex operator, and NADH but not NAD(+) inhibits T-Rex/DNA binding activity. A mechanism for redox sensing by Rex family members is proposed by analogy with domain closure of NAD(H)-dependent enzymes.
- Department of Biochemistry and Molecular Biology, Johns Hopkins Bloomberg School of Public Health, Baltimore, Maryland 21205, USA.
Organizational Affiliation: