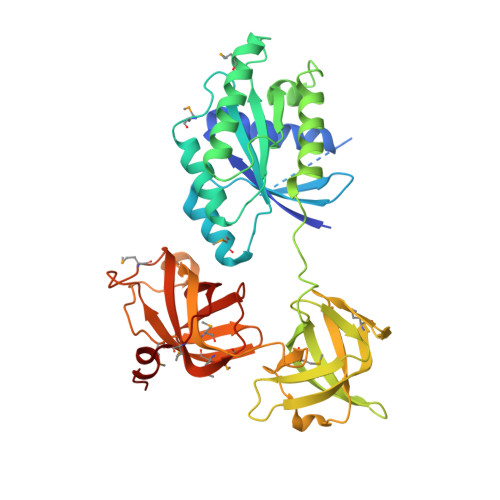
Crystal Structure of the Bovine Mitochondrial Elongation Factor Tu.Ts Complex
Jeppesen, M.G., Navratil, T., Spremulli, L.L., Nyborg, J.(2005) J Biological Chem 280: 5071-5081
- PubMed: 15557323 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411782200
- Primary Citation Related Structures:
1XB2 - PubMed Abstract:
The three-dimensional structure of the bovine mitochondrial elongation factor (EF)-Tu.Ts complex (EF-Tumt.Tsmt) has been determined to 2.2-A resolution using the multi-wavelength anomalous dispersion experimental method. This complex provides the first insight into the structure of EF-Tsmt. EF-Tsmt is similar to Escherichia coli and Thermus thermophilus EF-Ts in the amino-terminal domain. However, the structure of EF-Tsmt deviates considerably in the core domain with a five-stranded beta-sheet forming a portion of subdomain N of the core. In E. coli EF-Ts, this region is composed of a three-stranded sheet. The coiled-coil domain of the E. coli EF-Ts is largely eroded in EF-Tsmt, in which it consists of a large loop packed against subdomain C of the core. The conformation of bovine EF-Tumt in complex with EF-Tsmt is distinct from its conformation in the EF-Tumt.GDP complex. When domain III of bovine EF-Tumt.GDP is superimposed on domain III of EF-Tumt in the EF-Tumt.Tsmt complex, helix B from domain I is also almost superimposed. However, the rest of domain I is rotated relative to this helix toward domain II, which itself is rotated toward domain I relative to domain III. Extensive contacts are observed between the amino-terminal domain of EF-Tsmt and domain I of EF-Tumt. Furthermore, the conserved TDFV sequence of EF-Tsmt also contacts domain I with the side chain of Asp139 contacting helix B of EF-Tumt and inserting the side chain of Phe140 between helices B and C. The structure of the EF-Tumt.Tsmt complex provides new insights into the nucleotide exchange mechanism and provides a framework for explaining much of the mutational data obtained for this complex.
- Department of Molecular Biology, University of Aarhus, Gustav Wieds Vej 10 C, 8000 Aarhus C, Denmark.
Organizational Affiliation: