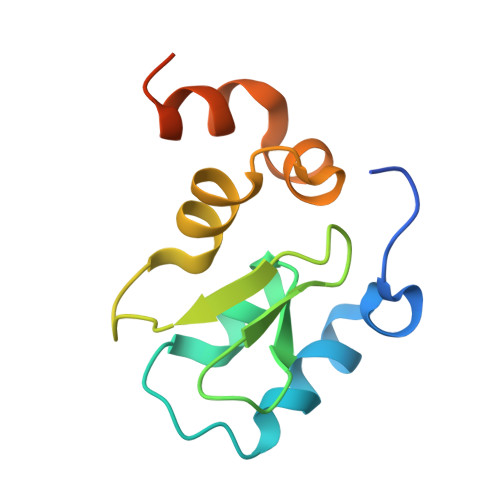
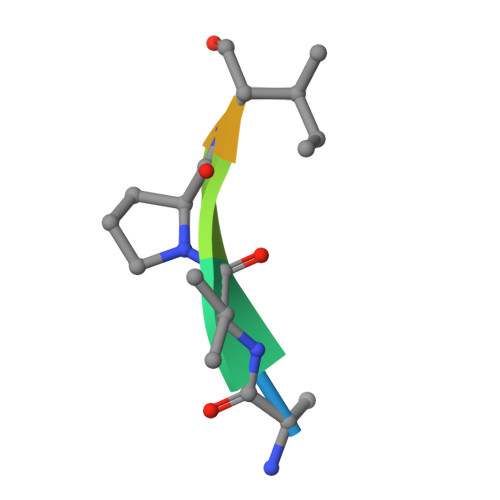
The BIR domain of IAP-like protein 2 is conformationally unstable: implications for caspase inhibition
Shin, H., Renatus, M., Eckelman, B.P., Nunes, V.A., Sampaio, C.A.M., Salvesen, G.S.(2005) Biochem J 385: 1-10
- PubMed: 15485395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20041107
- Primary Citation Related Structures:
1XB0, 1XB1 - PubMed Abstract:
Several IAP (inhibitor of apoptosis) proteins regulate cell fate decisions, and the X-linked IAP (XIAP) does so in part by inhibiting caspases, proteases that execute the apoptotic pathway. A tissue-specific homologue of XIAP, known as ILP2 (IAP-like protein 2), has previously been implicated in the control of apoptosis in the testis by direct inhibition of caspase 9. In examining this protein we found that the putative caspase 9 interaction domain is a surprisingly weak inhibitor and is also conformationally unstable. Comparison with the equivalent domain in XIAP demonstrated that the instability is due to the lack of a linker segment N-terminal to the inhibitory BIR (baculovirus IAP repeat) domain. Fusion of a 9-residue linker from XIAP to the N-terminus of ILP2 restored tight caspase 9 inhibition, dramatically increased conformational stability and allowed crystallization of the ILP2 BIR domain in a form strikingly similar to the XIAP third BIR domain. We conclude that ILP2 is an unstable protein, and cannot inhibit caspase 9 in a physiological way on its own. We speculate that ILP2 requires assistance from unidentified cellular factors to be an effective inhibitor of apoptosis in vivo.
- Program in Apoptosis and Cell Death Research, The Burnham Institute, 10901 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: