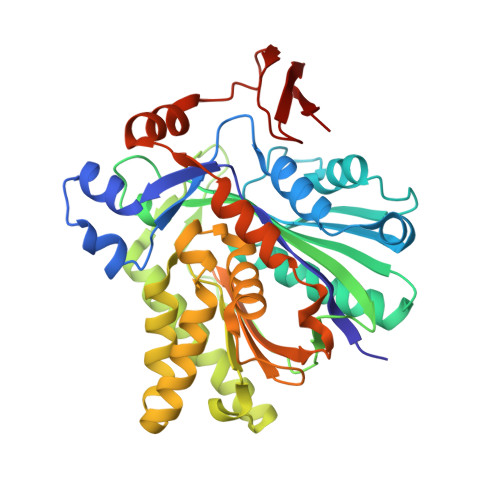
X-ray Crystal Structures of HMG-CoA Synthase from Enterococcus faecalis and a Complex with Its Second Substrate/Inhibitor Acetoacetyl-CoA
Steussy, C.N., Vartia, A.A., Burgner II, J.W., Sutherlin, A., Rodwell, V.W., Stauffacher, C.V.(2005) Biochemistry 44: 14256-14267
- PubMed: 16245942 Search on PubMed
- DOI: https://doi.org/10.1021/bi051487x
- Primary Citation Related Structures:
1X9E, 1YSL - PubMed Abstract:
Biosynthesis of the isoprenoid precursor, isopentenyl diphosphate, is a critical function in all independently living organisms. There are two major pathways for this synthesis, the non-mevalonate pathway found in most eubacteria and the mevalonate pathway found in animal cells and a number of pathogenic bacteria. An early step in this pathway is the condensation of acetyl-CoA and acetoacetyl-CoA into HMG-CoA, catalyzed by the enzyme HMG-CoA synthase. To explore the possibility of a small molecule inhibitor of the enzyme functioning as a non-cell wall antibiotic, the structure of HMG-CoA synthase from Enterococcus faecalis (MVAS) was determined by selenomethionine MAD phasing to 2.4 A and the enzyme complexed with its second substrate, acetoacetyl-CoA, to 1.9 A. These structures show that HMG-CoA synthase from Enterococcus is a member of the family of thiolase fold enzymes and, while similar to the recently published HMG-CoA synthase structures from Staphylococcus aureus, exhibit significant differences in the structure of the C-terminal domain. The acetoacetyl-CoA binary structure demonstrates reduced coenzyme A and acetoacetate covalently bound to the active site cysteine through a thioester bond. This is consistent with the kinetics of the reaction that have shown acetoacetyl-CoA to be a potent inhibitor of the overall reaction, and provides a starting point in the search for a small molecule inhibitor.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA.
Organizational Affiliation: