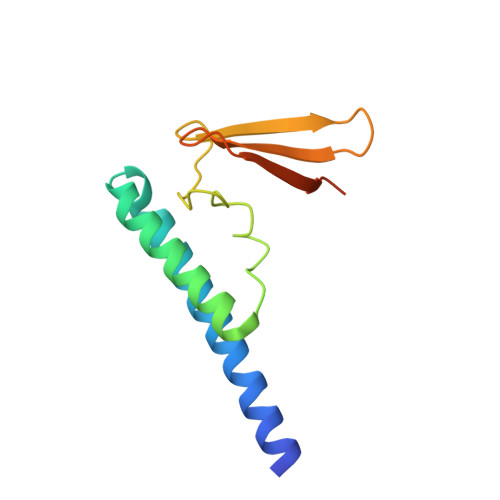
Crystal Structure of a Ribonuclease P Protein Ph1601p from Pyrococcus horikoshii OT3: An Archaeal Homologue of Human Nuclear Ribonuclease P Protein Rpp21(,)
Kakuta, Y., Ishimatsu, I., Numata, T., Kimura, K., Yao, M., Tanaka, I., Kimura, M.(2005) Biochemistry 44: 12086-12093
- PubMed: 16142906 Search on PubMed
- DOI: https://doi.org/10.1021/bi050738z
- Primary Citation Related Structures:
1X0T - PubMed Abstract:
Ribonuclease P (RNase P) is a ribonucleoprotein complex involved in the removal of 5' leader sequences from tRNA precursors (pre-tRNA). The human protein Rpp21 is essential for human RNase P activity in tRNA processing in vitro. The crystal structure of Ph1601p from the hyperthermophilic archaeon Pyrococcus horikoshii OT3, the archaeal homologue of Rpp21, was determined using the multiple anomalous dispersion (MAD) method with the aid of anomalous scattering in zinc and selenium at 1.6 A resolution. Ph1601p comprises an N-terminal domain (residues 1-55), a central linker domain (residues 56-79), and a C-terminal domain (residues 80-120), forming an L-shaped structure. The N-terminal domain consists of two long alpha-helices, while the central and C-terminal domains fold in a zinc ribbon domain. The electrostatic potential representation indicates the presence of positively charged clusters along the L arms, suggesting a possible role in RNA binding. A single zinc ion binds the well-ordered binding site that consists of four Cys residues (Cys68, Cys71, Cys97, and Cys100) and appears to stabilize the relative positions of the N- and C-domains. Mutations of Cys68 and Cys71 or Cys97 and Cys100 to Ser destabilize the protein structure, which results in inactivation of the RNase P activity. In addition, site-directed mutagenesis suggests that Lys69 at the central loop and Arg86 and Arg105 at the zinc ribbon domain are strongly involved in the functional activity, while Arg22, Tyr44, Arg65, and Arg84 play a modest role in the activity.
- Laboratory of Biochemistry, Department of Bioscience and Biotechnology, Faculty of Agriculture, Graduate School, Kyushu University, Fukuoka 812-8581, Japan.
Organizational Affiliation: