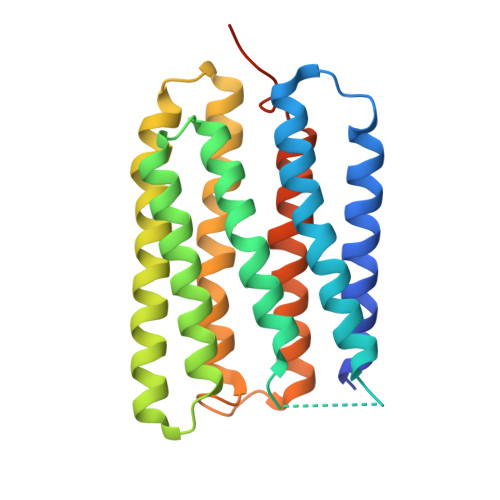
Crystal Structures of Acid Blue and Alkaline Purple Forms of Bacteriorhodopsin
Okumura, H., Murakami, M., Kouyama, T.(2005) J Mol Biology 351: 481-495
- PubMed: 16023672 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.06.026
- Primary Citation Related Structures:
1X0I, 1X0K - PubMed Abstract:
Bacteriorhodopsin, a light-driven proton pump found in the purple membrane of Halobacterium salinarum, exhibits purple at neutral pH but its color is sensitive to pH. Here, structures are reported for an acid blue form and an alkaline purple form of wild-type bacteriorhodopsin. When the P622 crystal prepared at pH 5.2 was acidified with sulfuric acid, its color turned to blue with a pKa of 3.5 and a Hill coefficient of 2. Diffraction data at pH 2-5 indicated that the purple-to-blue transition accompanies a large structural change in the proton release channel; i.e. the extracellular half of helix C moves towards helix G, narrowing the proton release channel and expelling a water molecule from a micro-cavity in the vicinity of the retinal Schiff base. In this respect, the acid-induced structural change resembles the structural change observed upon formation of the M intermediate. But, the acid blue form contains a sulfate ion in a site(s) near Arg82 that is created by re-orientations of the carboxyl groups of Glu194 and Glu204, residues comprising the proton release complex. This result suggests that proton uptake by the proton release complex evokes the anion binding, which in turn induces protonation of Asp85, a key residue regulating the absorption spectrum of the chromophore. Interestingly, a pronounced structural change in the proton release complex was also observed at high pH; i.e. re-orientation of Glu194 towards Tyr83 was found to take place at around pH 10. This alkaline transition is suggested to be accompanied by proton release from the proton release complex and responsible for rapid formation of the M intermediate at high pH.
- Department of Physics, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan.
Organizational Affiliation: