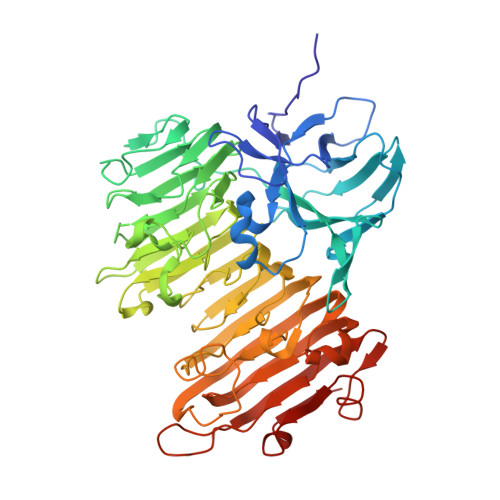
Crystal Structure of Aspergillus niger Isopullulanase, a Member of Glycoside Hydrolase Family 49
Mizuno, M., Koide, A., Yamamura, A., Akeboshi, H., Yoshida, H., Kamitori, S., Sakano, Y., Nishikawa, A., Tonozuka, T.(2008) J Mol Biology 376: 210-220
- PubMed: 18155243 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.11.098
- Primary Citation Related Structures:
1WMR, 1X0C, 2Z8G - PubMed Abstract:
An isopullulanase (IPU) from Aspergillus niger ATCC9642 hydrolyzes alpha-1,4-glucosidic linkages of pullulan to produce isopanose. Although IPU does not hydrolyze dextran, it is classified into glycoside hydrolase family 49 (GH49), major members of which are dextran-hydrolyzing enzymes. IPU is highly glycosylated, making it difficult to obtain its crystal. We used endoglycosidase H(f) to cleave the N-linked oligosaccharides of IPU, and we here determined the unliganded and isopanose-complexed forms of IPU, both solved at 1.7-A resolution. IPU is composed of domains N and C joined by a short linker, with electron density maps for 11 or 12 N-acetylglucosamine residues per molecule. Domain N consists of 13 beta-strands and forms a beta-sandwich. Domain C, where the active site is located, forms a right-handed beta-helix, and the lengths of the pitches of each coil of the beta-helix are similar to those of GH49 dextranase and GH28 polygalacturonase. The entire structure of IPU resembles that of a GH49 enzyme, Penicillium minioluteum dextranase (Dex49A), despite a difference in substrate specificity. Compared with the active sites of IPU and Dex49A, the amino acid residues participating in subsites +2 and +3 are not conserved, and the glucose residues of isopanose bound to IPU completely differ in orientation from the corresponding glucose residues of isomaltose bound to Dex49A. The shape of the catalytic cleft characterized by the seventh coil of the beta-helix and a loop from domain N appears to be critical in determining the specificity of IPU for pullulan.
- Department of Applied Biological Science, Tokyo University of Agriculture and Technology, 3-5-8 Saiwai-Cho, Fuchu, Tokyo 183-8509, Japan.
Organizational Affiliation: