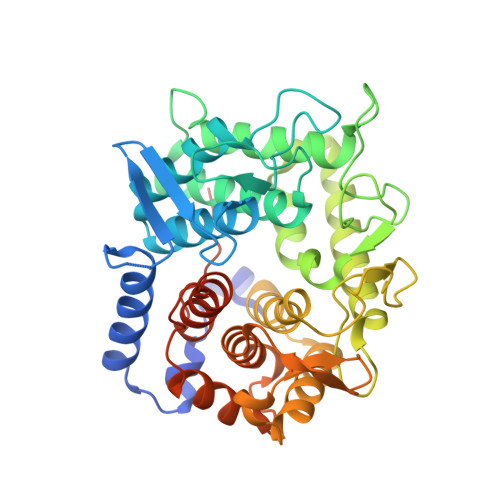
Structural Basis for the Specificity of the Reducing End Xylose-releasing Exo-oligoxylanase from Bacillus halodurans C-125
Fushinobu, S., Hidaka, M., Honda, Y., Wakagi, T., Shoun, H., Kitaoka, M.(2005) J Biological Chem 280: 17180-17186
- PubMed: 15718242
- DOI: https://doi.org/10.1074/jbc.M413693200
- Primary Citation Related Structures:
1WU4, 1WU5, 1WU6 - PubMed Abstract:
Reducing end xylose-releasing exo-oligoxylanase from Bacillus halodurans C-125 (Rex) hydrolyzes xylooligosaccharides whose degree of polymerization is greater than or equal to 3, releasing the xylose unit at the reducing end. It is a unique exo-type glycoside hydrolase that recognizes the xylose unit at the reducing end in a very strict manner, even discriminating the beta-anomeric hydroxyl configuration from the alpha-anomer or 1-deoxyxylose. We have determined the crystal structures of Rex in unliganded and complex forms at 1.35-2.20-A resolution and revealed the structural aspects of its three subsites ranging from -2 to +1. The structure of Rex was compared with those of endo-type enzymes in glycoside hydrolase subfamily 8a (GH-8a). The catalytic machinery of Rex is basically conserved with other GH-8a enzymes. However, subsite +2 is blocked by a barrier formed by a kink in the loop before helix alpha10. His-319 in this loop forms a direct hydrogen bond with the beta-hydroxyl of xylose at subsite +1, contributing to the specific recognition of anomers at the reducing end.
- Department of Biotechnology, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan. asfushi@mail.ecc.u-tokyo.ac.jp
Organizational Affiliation: