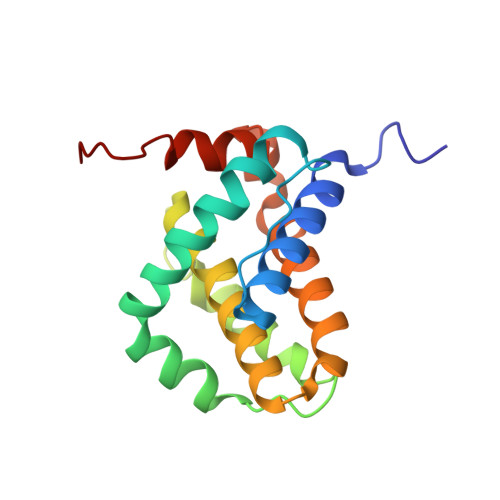
Solution Structure of Prosurvival Mcl-1 and Characterization of Its Binding by Proapoptotic BH3-only Ligands
Day, C.L., Chen, L., Richardson, S.J., Harrison, P.J., Huang, D.C., Hinds, M.G.(2005) J Biological Chem 280: 4738-4744
- PubMed: 15550399
- DOI: https://doi.org/10.1074/jbc.M411434200
- Primary Citation of Related Structures:
1WSX - PubMed Abstract:
The B cell lymphoma-2 (Bcl-2) homologs myeloid cell leukemia-1 (Mcl-1) and A1 are prosurvival factors that selectively bind a subset of proapoptotic Bcl homology (BH) 3-only proteins. To investigate the molecular basis of the selectivity, we determined the solution structure of the C-terminal Bcl-2-like domain of Mcl-1. This domain shares features expected of a prosurvival Bcl-2 protein, having a helical fold centered on a core hydrophobic helix and a surface-exposed hydrophobic groove for binding its cognate partners. A number of residues in the binding groove differentiate Mcl-1 from its homologs, and in contrast to other Bcl-2 homologs, Mcl-1 has a binding groove in a conformation intermediate between the open structures characterized by peptide complexes and the closed state observed in unliganded structures. Mutagenesis of potential binding site residues was used to probe the contributions of groove residues to the binding properties of Mcl-1. Although mutations in Mcl-1 had little impact on binding, a single mutation in the BH3-only ligand Bad enabled it to bind both Mcl-1 and A1 while retaining its binding to Bcl-2, Bcl-xL, and Bcl-w. Elucidating the selective action of certain BH3-only ligands is required for delineating their mode of action and will aid the search for effective BH3-mimetic drugs.
- Department of Biochemistry, University of Otago, Dunedin 9001, New Zealand.
Organizational Affiliation: