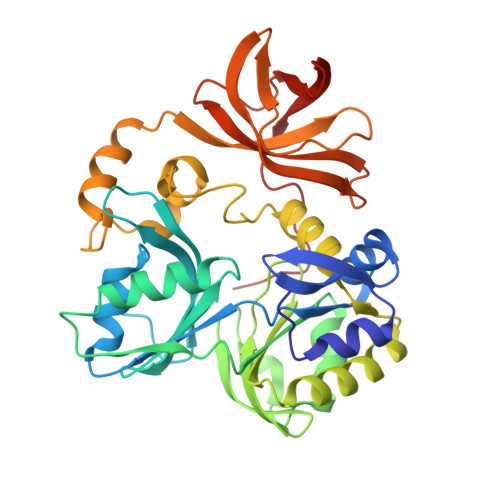
Crystal Structure of T-protein of the Glycine Cleavage System: Cofactor binding, insights into H-protein recognition, and molecular basis for understanding nonketotic hyperglycinemia
Lee, H.H., Kim, D.J., Ahn, H.J., Ha, J.Y., Suh, S.W.(2004) J Biological Chem 279: 50514-50523
- PubMed: 15355973 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409672200
- Primary Citation Related Structures:
1WOO, 1WOP, 1WOR, 1WOS - PubMed Abstract:
The glycine cleavage system catalyzes the oxidative decarboxylation of glycine in bacteria and in mitochondria of animals and plants. Its deficiency in human causes nonketotic hyperglycinemia, an inborn error of glycine metabolism. T-protein, one of the four components of the glycine cleavage system,is a tetrahydrofolate dependent aminomethyltransferase. It catalyzes the transfer of the methylene carbon unit to tetrahydrofolate from the methylamine group covalently attached to the lipoamide arm of H-protein. To gain insight into the T-protein function at the molecular level, we have determined the first crystal structure of T-protein from Thermotoga maritima by the multiwavelength anomalous diffraction method of x-ray crystallography and refined four structures: the apoform; the tetrahydrofolate complex; the folinic acid complex; and the lipoic acid complex. The overall fold of T-protein is similar to that of the C-terminal tetrahydrofolate-binding region (residues 421-830) of Arthrobacter globiformis dimethylglycine oxidase. Tetrahydrofolate (or folinic acid) is bound near the center of the tripartite T-protein. Lipoic acid is bound adjacent to the tetrahydrofolate binding pocket, thus defining the interaction surface for H-protein binding. A homology model of the human T-protein provides the structural framework for understanding the molecular mechanisms underlying the development of nonketotic hyperglycinemia due to missense mutations of the human T-protein.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: