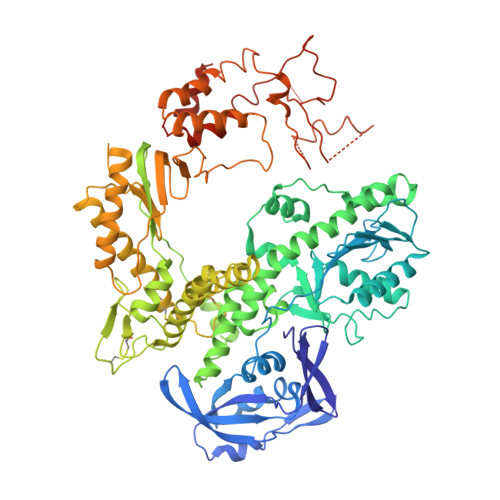
Crystal Structure of DNA Polymerase from Hyperthermophilic Archaeon Pyrococcus Kodakaraensis Kod1
Hashimoto, H., Nishioka, M., Fujiwara, S., Takagi, M., Imanaka, T., Inoue, T., Kai, Y.(2001) J Mol Biology 306: 469-477
- PubMed: 11178906
- DOI: https://doi.org/10.1006/jmbi.2000.4403
- Primary Citation of Related Structures:
1WNS - PubMed Abstract:
The crystal structure of family B DNA polymerase from the hyperthermophilic archaeon Pyrococcus kodakaraensis KOD1 (KOD DNA polymerase) was determined. KOD DNA polymerase exhibits the highest known extension rate, processivity and fidelity. We carried out the structural analysis of KOD DNA polymerase in order to clarify the mechanisms of those enzymatic features. Structural comparison of DNA polymerases from hyperthermophilic archaea highlighted the conformational difference in Thumb domains. The Thumb domain of KOD DNA polymerase shows an "opened" conformation. The fingers subdomain possessed many basic residues at the side of the polymerase active site. The residues are considered to be accessible to the incoming dNTP by electrostatic interaction. A beta-hairpin motif (residues 242-249) extends from the Exonuclease (Exo) domain as seen in the editing complex of the RB69 DNA polymerase from bacteriophage RB69. Many arginine residues are located at the forked-point (the junction of the template-binding and editing clefts) of KOD DNA polymerase, suggesting that the basic environment is suitable for partitioning of the primer and template DNA duplex and for stabilizing the partially melted DNA structure in the high-temperature environments. The stabilization of the melted DNA structure at the forked-point may be correlated with the high PCR performance of KOD DNA polymerase, which is due to low error rate, high elongation rate and processivity.
- Department of Materials Chemistry, Graduate School of Engineering, Osaka University, Suita, Japan.
Organizational Affiliation: