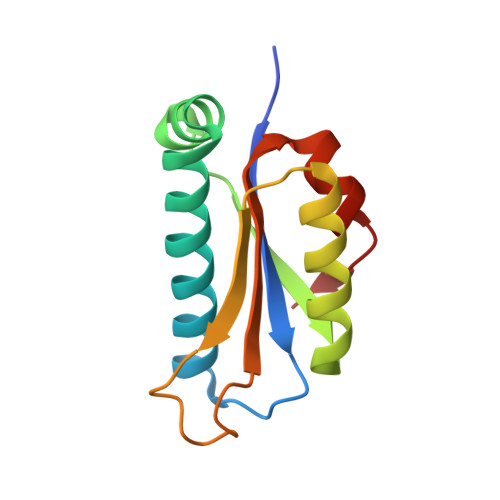
Structure of peptidyl-tRNA hydrolase 2 from Pyrococcus horikoshii OT3: insight into the functional role of its dimeric state.
Shimizu, K., Kuroishi, C., Sugahara, M., Kunishima, N.(2008) Acta Crystallogr D Biol Crystallogr 64: 444-453
- PubMed: 18391411 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908002850
- Primary Citation Related Structures:
1WN2, 2D3K - PubMed Abstract:
Peptidyl-tRNA hydrolases catalyze the hydrolytic removal of the peptidyl moiety from the peptidyl-tRNA molecule to prevent misreading during translation. Here, the expression, purification, crystallization and X-ray diffraction study of peptidyl-tRNA hydrolase 2 from Pyrococcus horikoshii OT3 (PhPth2) are described. The crystal structures were determined as similar biological dimers in two different forms: P4(1)2(1)2 at 1.2 A resolution (form 1) and P4(3)22 at 1.9 A resolution (form 2). In the form 1 structure, the asymmetric unit contains one PhPth2 subunit and a crystallographic twofold axis defines the dimeric association with the cognate subunit. In the form 2 structure, there are two PhPth2 subunits in the asymmetric unit that make a similar dimer with a noncrystallographic twofold axis. In order to evaluate the thermodynamic stability, the intra-protomer and inter-protomer interactions of PhPth2 were analyzed and compared with those of other Pth2-family members. The thermodynamic parameters show that the large number of ion pairs compared with family members from other mesophilic organisms would contribute to the thermostability of PhPth2. The structural difference between the two dimers was quantitatively evaluated by a multiple C(alpha)-atom superposition. A significant structural difference between the two dimers was observed around the putative active site of this enzyme. A rigid-body rotation takes place so as to retain the dimeric twofold symmetry, suggesting positive cooperativity upon tRNA binding. The mechanism of ligand binding was further investigated using a docking model with a tRNA molecule. The docking study suggests that the binding of tRNA requires its simultaneous interaction with both subunits of the PhPth2 dimer.
- Advanced Protein Crystallography Research Group, RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: