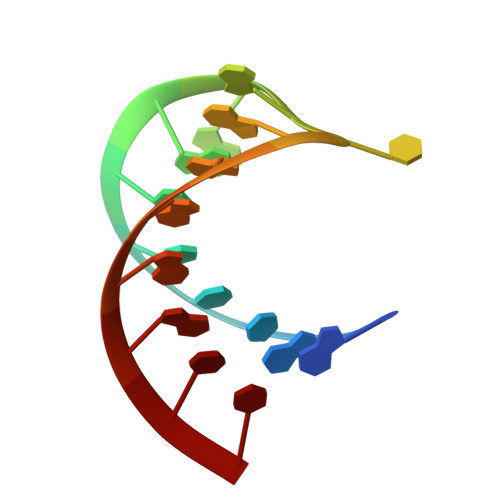
Solution structure of an RNA stem-loop derived from the 3' conserved region of eel LINE UnaL2
Baba, S., Kajikawa, M., Okada, N., Kawai, G.(2004) RNA 10: 1380-1387
- PubMed: 15273327
- DOI: https://doi.org/10.1261/rna.7460104
- Primary Citation Related Structures:
1WKS - PubMed Abstract:
The eel long interspersed element (LINE) UnaL2 and its partner short interspersed element (SINE) share a conserved 3' tail containing a stem-loop that is critical for their retrotransposition. Presumably, the first step of retrotransposition is the recognition of their 3' tails by UnaL2-encoded reverse transcriptase. The solution structure of a 17-nucleotide RNA derived from the 3' tail of UnaL2 was determined by NMR. The GGAUA loop forms a specific structure in which the uridine is exposed to solvent with the third and fifth adenosines stacked. A sharp turn in the phosphodiester backbone occurs between the second guanosine and third adenosine. When the uridine is mutated (but not deleted), all mutants form the loop structure, indicating that the loop structure requires an exposed fourth residue. The retrotransposition assay in HeLa cells revealed that retrotransposition requires the second guanosine, although any nucleoside functions at the fourth position, suggesting that UnaL2 reverse transcriptase specifically recognizes the 5' side of the GGANA loop.
- Department of Industrial Chemistry, Faculty of Engineering, Chiba Institute of Technology, 2-17-1 Tsudanuma, Narashino, 275-0016, Japan.
Organizational Affiliation: