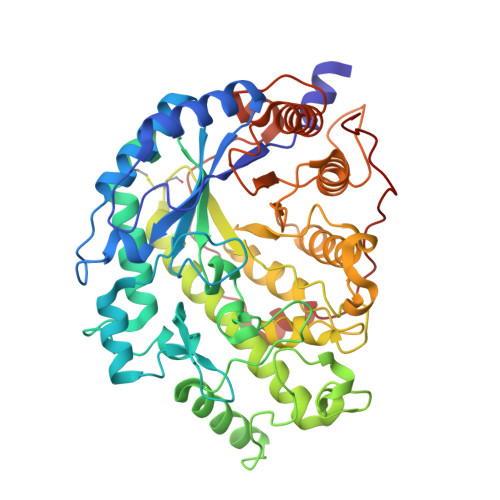
Structural analysis of threonine 342 mutants of soybean beta-amylase: role of a conformational change of the inner loop in the catalytic mechanism.
Kang, Y.N., Tanabe, A., Adachi, M., Utsumi, S., Mikami, B.(2005) Biochemistry 44: 5106-5116
- PubMed: 15794648 Search on PubMed
- DOI: https://doi.org/10.1021/bi0476580
- Primary Citation Related Structures:
1WDP, 1WDQ, 1WDR, 1WDS - PubMed Abstract:
Two different conformations of the inner loop (residues 340-346) have been found in the soybean beta-amylase structures. In the "product form", the Thr 342 residue creates hydrogen bonds with Glu 186 (catalytic acid) and with the glucose residues at subsites -1 and +1, whereas most of those interactions are lost in the "apo form". To elucidate the relationship between the structural states of the inner loop and the catalytic mechanism, Thr 342 was mutated to Val, Ser, and Ala, respectively, and their crystal structures complexed with maltose were determined together with that of the apo enzyme at 1.27-1.64 A resolutions. The k(cat) values of the T342V, T342S, and T342A mutants decreased by 13-, 360-, and 1700-fold, respectively, compared to that of the wild-type enzyme. Whereas the inner loops in the wild-type/maltose and T342V/maltose complexes adopted the product form, those of the T342S/maltose and T342A/maltose complexes showed the apo form. Structural analyses suggested that the side chain of Thr 342 in product form plays an important role in distorting the sugar ring at subsite -1, stabilizing the deprotonated form of Glu 186, and grasping the glucose residue of the remaining substrate at subsite +1. The third hypothesis was proved by the fact that T342V hydrolyzes maltoheptaose following only multichain attack in contrast to multiple attack of the wild-type enzyme.
- Laboratory of Food Quality Design and Development, Graduate School of Agriculture, Kyoto University, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: