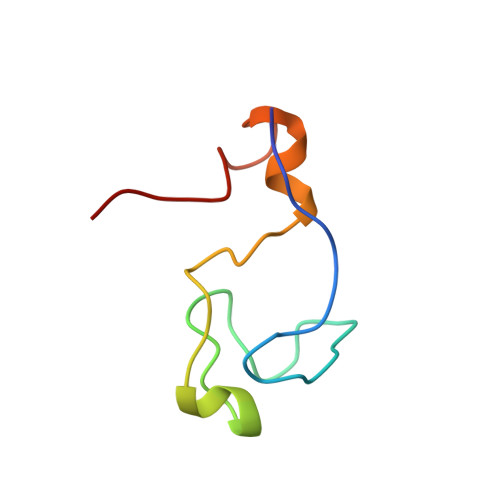
Structure of the C-terminal RING Finger from a RING-IBR-RING/TRIAD Motif Reveals a Novel Zinc-binding Domain Distinct from a RING
Capili, A.D., Edghill, E.L., Wu, K., Borden, K.L.B.(2004) J Mol Biology 340: 1117-1129
- PubMed: 15236971
- DOI: https://doi.org/10.1016/j.jmb.2004.05.035
- Primary Citation of Related Structures:
1WD2 - PubMed Abstract:
The really interesting new gene (RING) family of proteins contains over 400 members with diverse physiological functions. A subset of these domains is found in the context of the RING-IBR-RING/TRIAD motifs which function as E3 ubiquitin ligases. Our sequence analysis of the C-terminal RING (RING2) from this motif show that several metal ligating and hydrophobic residues critical for the formation of a classical RING cross-brace structure are not present. Thus, we determined the structure of the RING2 from the RING-IBR-RING motif of HHARI and showed that RING2 has a completely distinct topology from classical RINGs. Notably, RING2 binds only one zinc atom per monomer rather than two and uses a different hydrophobic network to that of classical RINGs. Additionally, this RING2 topology is novel, bearing slight resemblance to zinc-ribbon motifs around the zinc site and is different from the topologies of the zinc binding sites found in RING and PHDs. We demonstrate that RING2 acts as an E3 ligase in vitro and using mutational analysis deduce the structural features required for this activity. Further, mutations in the RING-IBR-RING of Parkin cause a rare form of Parkinsonism and these studies provide an explanation for those mutations that occur in its RING2. From a comparison of the RING2 structure with those reported for RINGs, we infer sequence determinants that allow discrimination between RING2 and RING domains at the sequence analysis level.
- Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, New York University, New York, NY 10029, USA.
Organizational Affiliation: