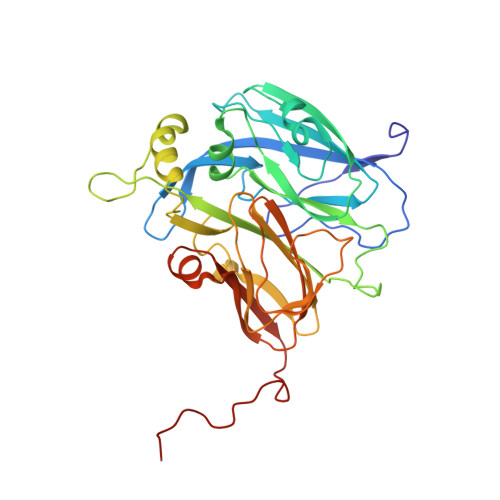
Observation of an Unprecedented Cu Bis-His Site: Crystal Structure of the H129V Mutant of Nitrite Reductase
Ellis, M.J., Antonyuk, S.V., Strange, R.W., Sawers, G., Eady, R.R., Hasnain, S.S.(2004) Inorg Chem 43: 7591
- PubMed: 15554622 Search on PubMed
- DOI: https://doi.org/10.1021/ic048966p
- Primary Citation Related Structures:
1WAE - PubMed Abstract:
Copper nitrite reductases contain both an electron-transfer type 1 Cu site and a catalytic type 2 Cu site. We have mutated one of the type 2 copper ligating histidines to observe the effect on catalytic turnover. This mutation has created a unique site where Cu is ligated by 2 His Nepsilon2 atoms alone.
- Molecular Biophysics Group, CCLRC Daresbury Laboratory, Warrington, Cheshire, UK.
Organizational Affiliation: