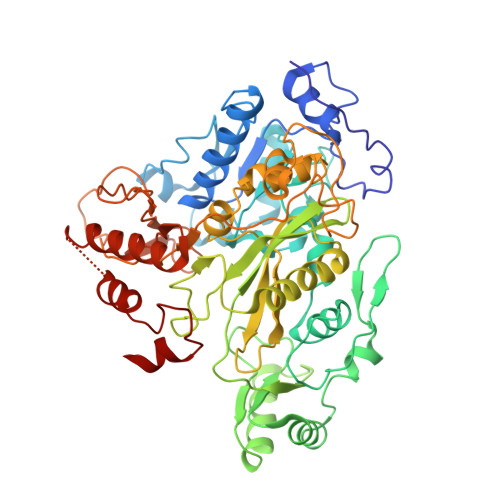
A Mechanism for the Potent Inhibition of Eukaryotic Acetyl-Coenzyme a Carboxylase by Soraphen A, a Macrocyclic Polyketide Natural Product
Shen, Y., Volrath, S.L., Weatherly, S.C., Elich, T.D., Tong, L.(2004) Mol Cell 16: 881
- PubMed: 15610732 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.11.034
- Primary Citation Related Structures:
1W93, 1W96 - PubMed Abstract:
Acetyl-coenzyme A carboxylases (ACCs) have crucial roles in fatty acid metabolism. Soraphen A, a macrocyclic polyketide natural product, is a nanomolar inhibitor against the biotin carboxylase (BC) domain of human, yeast, and other eukaryotic ACCs. Here we report the crystal structures of the yeast BC domain, alone and in complex with soraphen A. Soraphen has extensive interactions with an allosteric site, about 25 A from the active site. The specificity of soraphen is explained by large structural differences between the eukaryotic and prokaryotic BC in its binding site, confirmed by our studies on the effects of single-site mutations in this binding site. Unexpectedly, our structures suggest that soraphen may bind in the BC dimer interface and inhibit the BC activity by disrupting the oligomerization of this domain. Observations from native gel electrophoresis confirm this structural insight. The structural information provides a foundation for structure-based design of new inhibitors against these enzymes.
- Department of Biological Sciences, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: