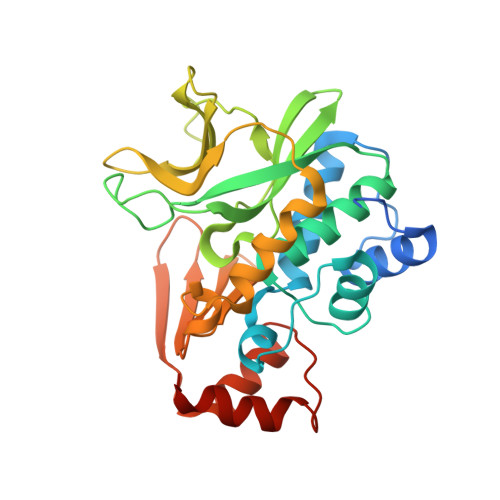
Expression, purification, characterization and structure of Pseudomonas aeruginosa arylamine N-acetyltransferase.
Westwood, I.M., Holton, S.J., Rodrigues-Lima, F., Dupret, J.M., Bhakta, S., Noble, M.E., Sim, E.(2005) Biochem J 385: 605-612
- PubMed: 15447630 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20041330
- Primary Citation Related Structures:
1W4T - PubMed Abstract:
The gene for NAT (arylamine N-acetyltransferase) from Pseudomonas aeruginosa (panat) has been cloned from genomic DNA, and the gene product (PANAT) expressed as an N-terminal histidine-tagged protein in Escherichia coli and purified via nickel ion affinity chromatography. The specific activities of PANAT against a broad range of substrates have been investigated and compared with those of other prokaryotic NAT enzymes. For most arylamine substrates identified, PANAT exhibits in vitro specific activities typically one order of magnitude greater than those of recombinant NAT enzymes from Mycobacterium smegmatis or Salmonella typhimurium. Among the substrates of PANAT so far identified are the anti-tubercular drug isoniazid, 5-aminosalicylate (a drug used in the treatment of inflammatory bowel disease), as well as important environmental pollutants such as 3,4-dichloroaniline and 2-aminofluorene. As well as acetylating common NAT substrates, PANAT is unique among the prokaryotic NATs so far studied in acetylating the folate precursor 4-aminobenzoic acid and the folate catabolite 4-aminobenzoylglutamate. The recombinant protein has been expressed in sufficient quantity to allow protein crystallization, and we have subsequently determined the 1.95 A structure of PANAT by X-ray crystallography.
- Department of Pharmacology, University of Oxford, Mansfield Road, Oxford OX1 3QT, UK.
Organizational Affiliation: