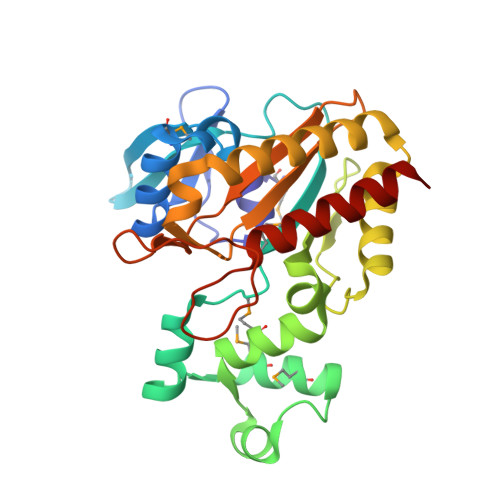
Structure of a Human Inositol 1,4,5-Trisphosphate 3-Kinase; Substrate Binding Reveals Why It is not a Phosphoinositide 3-Kinase
Gonzalez, B., Schell, M.J., Letcher, A.J., Veprintsev, D.B., Irvine, R.F., Williams, R.L.(2004) Mol Cell 15: 689
- PubMed: 15350214 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.08.004
- Primary Citation Related Structures:
1W2C, 1W2D, 1W2F - PubMed Abstract:
Mammalian cells produce a variety of inositol phosphates (InsPs), including Ins(1,4,5)P3 that serves both as a second messenger and as a substrate for inositol polyphosphate kinases (IPKs), which further phosphorylate it. We report the structure of an IPK, the human Ins(1,4,5)P3 3-kinase-A, both free and in complexes with substrates and products. This enzyme catalyzes transfer of a phosphate from ATP to the 3-OH of Ins(1,4,5)P3, and its X-ray crystal structure provides a template for understanding a broad family of InsP kinases. The catalytic domain consists of three lobes. The N and C lobes bind ATP and resemble protein and lipid kinases, despite insignificant sequence similarity. The third lobe binds inositol phosphate and is a unique four-helix insertion in the C lobe. This lobe embraces all of the phosphates of Ins(1,4,5)P3 in a positively charged pocket, explaining the enzyme's substrate specificity and its inability to phosphorylate PtdIns(4,5)P2, the membrane-resident analog of Ins(1,4,5)P3.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: