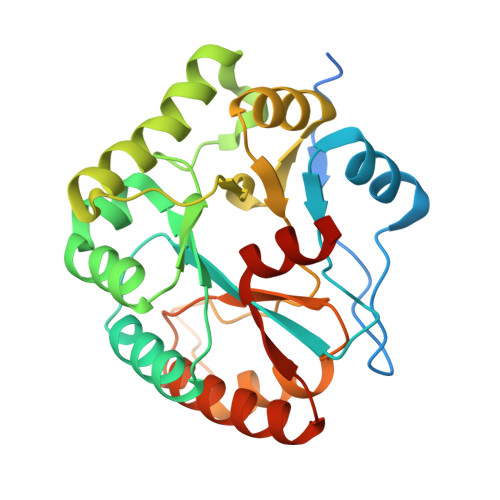
Structures of Bacillus subtilis PdaA, a family 4 carbohydrate esterase, and a complex with N-acetyl-glucosamine.
Blair, D.E., van Aalten, D.M.(2004) FEBS Lett 570: 13-19
- PubMed: 15251431 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2004.06.013
- Primary Citation Related Structures:
1W17, 1W1A, 1W1B - PubMed Abstract:
Family 4 carbohydrate esterases deacetylate polymeric carbohydrate substrates such as chitin, acetyl xylan and peptidoglycan. Although some of these enzymes have recently been enzymologically characterised, neither their structure nor their reaction mechanism has been defined. Sequence conservation in this family has pointed to a conserved core, termed the NodB homology domain. We describe the cloning, purification and 1.9 A crystal structure of PdaA, a peptidoglycan deacetylase from Bacillus subtilis. The enzyme assumes a fold related to a (beta/alpha)8 barrel, with a long groove on the surface of the protein that harbours all conserved residues. A complex with the substrate analogue N-acetyl-glucosamine was refined to 2.25 A resolution, revealing interactions of an aspartic acid and three histidines, all conserved in the NodB homology domain, with the ligand. The PdaA structure provides a template for interpreting the wealth of sequence data on family 4 carbohydrate esterases in a structural context and represents a first step towards understanding the reaction mechanism of this family of enzymes.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: