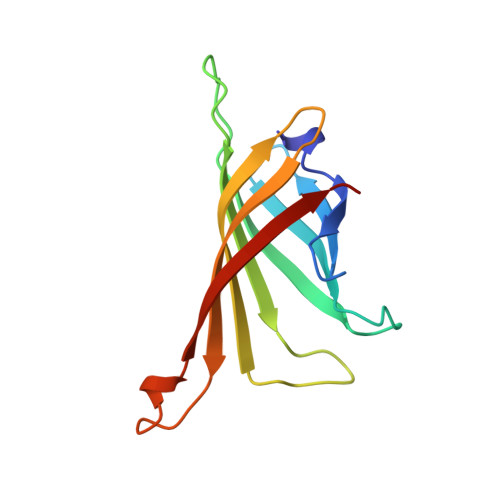
In crystals of complexes of streptavidin with peptide ligands containing the HPQ sequence the pKa of the peptide histidine is less than 3.0.
Katz, B.A., Cass, R.T.(1997) J Biological Chem 272: 13220-13228
- PubMed: 9148939 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.272.20.13220
- Primary Citation Related Structures:
1VWA, 1VWB, 1VWC, 1VWD, 1VWE, 1VWF, 1VWG, 1VWH, 1VWI, 1VWJ, 1VWK, 1VWL, 1VWM, 1VWN, 1VWO, 1VWP, 1VWQ, 1VWR - PubMed Abstract:
The pH dependences of the affinities for streptavidin of linear and cyclic peptide ligands containing the HPQ sequence discovered by phage display were determined by plasmon resonance measurements. At pH values ranging from 3.0 to 9.0, the Kd values for Ac-AEFSHPQNTIEGRK-NH2, cyclo-Ac-AE[CHPQGPPC]IEGRK-NH2, and cyclo-Ac-AE[CHPQFC]IEGRK-NH2, were determined by competition, and those for cyclo-[5-S-valeramide-HPQGPPC]K-NH2 were determined directly by equilibrium affinity measurements. The Kd values of the ligands increase by an average factor of 3.0 +/- 0.8 per decrease in pH unit between pH approximately 4.5 and pH approximately 6.3. Below pH approximately 4.5 there is a smaller increase in Kd values, and above pH approximately 6.3 the Kd values become relatively pH-independent. We determined the crystal structures of complexes of streptavidin with cyclo-[5-S-valeramide-HPQGPPC]K-NH2 at pH 1.5, 2.5, 3.0, and 3.5, with cyclo-Ac-[CHPQFC]-NH2 at pH 2.0, 3.0, 3.6, 4.2, 4.8, and 11.8, with cyclo-Ac-[CHPQGPPC]-NH2 at pH 2.5, 2.9, and 3.7, and with FSHPQNT at pH 4.0 and compared the structures with one another and with those previously determined at other pH values. At pH values from 3.0 to 11.8, the electron density for the peptide His side chain is strong, flat, and well defined. A hydrogen bond between the Ndelta1 atom of the His and the peptide Gln amide group indicates the His of the bound peptide in the crystals is uncharged at pH >/= 3.0. By determining selected structures in two different space groups, I222 with two crystallographically inequivalent ligand sites and I4122 with one site, we show that below pH approximately 3.0, the pKa of the bound peptide His in the crystals is influenced by crystal packing interactions. The presence of the Ndelta1His-NGln hydrogen bond along with pH dependences of the peptide affinities suggest that deprotonation of the peptide His is required for high affinity binding of HPQ-containing peptides to streptavidin both in the crystals and in solution.
- Arris Pharmaceutical Corporation, South San Francisco, California 94080, USA. bak@arris.com
Organizational Affiliation: