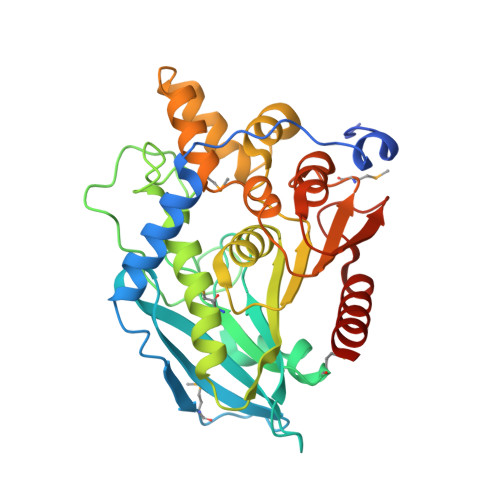
Functional and structural characterization of a thermostable acetyl esterase from Thermotoga maritima.
Levisson, M., Han, G.W., Deller, M.C., Xu, Q., Biely, P., Hendriks, S., Ten Eyck, L.F., Flensburg, C., Roversi, P., Miller, M.D., McMullan, D., von Delft, F., Kreusch, A., Deacon, A.M., van der Oost, J., Lesley, S.A., Elsliger, M.A., Kengen, S.W., Wilson, I.A.(2012) Proteins 80: 1545-1559
- PubMed: 22411095 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24041
- Primary Citation Related Structures:
1VLQ, 3M81, 3M82, 3M83 - PubMed Abstract:
TM0077 from Thermotoga maritima is a member of the carbohydrate esterase family 7 and is active on a variety of acetylated compounds, including cephalosporin C. TM0077 esterase activity is confined to short-chain acyl esters (C2-C3), and is optimal around 100°C and pH 7.5. The positional specificity of TM0077 was investigated using 4-nitrophenyl-β-D-xylopyranoside monoacetates as substrates in a β-xylosidase-coupled assay. TM0077 hydrolyzes acetate at positions 2, 3, and 4 with equal efficiency. No activity was detected on xylan or acetylated xylan, which implies that TM0077 is an acetyl esterase and not an acetyl xylan esterase as currently annotated. Selenomethionine-substituted and native structures of TM0077 were determined at 2.1 and 2.5 Å resolution, respectively, revealing a classic α/β-hydrolase fold. TM0077 assembles into a doughnut-shaped hexamer with small tunnels on either side leading to an inner cavity, which contains the six catalytic centers. Structures of TM0077 with covalently bound phenylmethylsulfonyl fluoride and paraoxon were determined to 2.4 and 2.1 Å, respectively, and confirmed that both inhibitors bind covalently to the catalytic serine (Ser188). Upon binding of inhibitor, the catalytic serine adopts an altered conformation, as observed in other esterase and lipases, and supports a previously proposed catalytic mechanism in which Ser hydroxyl rotation prevents reversal of the reaction and allows access of a water molecule for completion of the reaction.
- Laboratory of Microbiology, Wageningen University, Dreijenplein 10, 6703 HB, Wageningen, The Netherlands.
Organizational Affiliation: