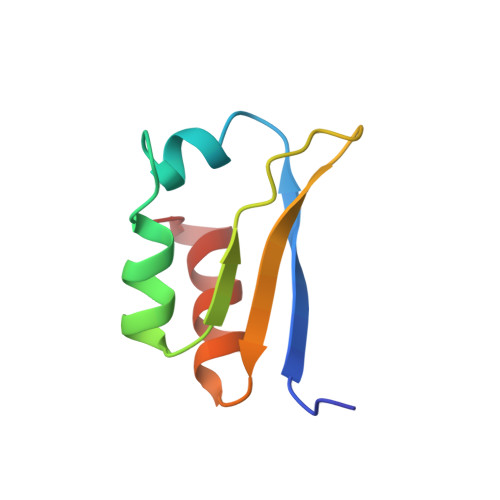
Three-dimensional structure and stability of the KH domain: molecular insights into the fragile X syndrome.
Musco, G., Stier, G., Joseph, C., Castiglione Morelli, M.A., Nilges, M., Gibson, T.J., Pastore, A.(1996) Cell 85: 237-245
- PubMed: 8612276 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81100-9
- Primary Citation Related Structures:
1VIG, 1VIH - PubMed Abstract:
The KH module is a sequence motif found in a number of proteins that are known to be in close association with RNA. Experimental evidence suggests a direct involvement of KH in RNA binding. The human FMR1 protein, which has two KH domains, is associated with fragile X syndrome, the most common inherited cause of mental retardation. Here we present the three-dimensional solution structure of the KH module. The domain consists of a stable beta alpha alpha beta beta alpha fold. On the basis of our results, we suggest a potential surface for RNA binding centered on the loop between the first two helices. Substitution of a well-conserved hydrophobic residue located on the second helix destroys the KH fold; a mutation of this position in FMR1 leads to an aggravated fragile X phenotype.
- European Molecular Biology Laboratory, Heidelberg, Federal Republic of Germany.
Organizational Affiliation: