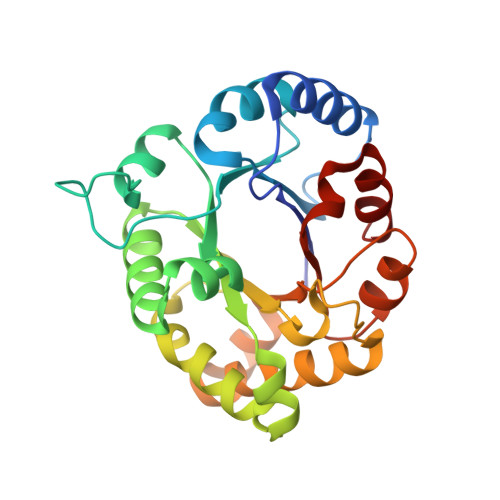
Structures of Unliganded and Inhibitor Complexes of W168F, a Loop6 Hinge Mutant of Plasmodium falciparum Triosephosphate Isomerase: Observation of an Intermediate Position of Loop6
Eaazhisai, K., Balaram, H., Balaram, P., Murthy, M.R.N.(2004) J Mol Biology 343: 671-684
- PubMed: 15465054 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.08.060
- Primary Citation Related Structures:
1VGA, 1WOA, 1WOB - PubMed Abstract:
The enzymatic reaction of triosephosphate isomerase (TIM) is controlled by the movement of a loop (loop6, residues 166-176). Crystal structures of TIMs from a variety of sources have revealed that the loop6, which is in an open conformation in the unliganded enzyme, adopts a closed conformation in inhibitor complexes. In contrast, structures with loop open conformation are obtained in most of the complexes of TIM from the malarial parasite Plasmodium falciparum (PfTIM). W168 is a conserved N-terminal hinge residue, involved in different sets of interactions in the "open" and "closed" forms of loop6. The role of W168 in determining the loop conformation was examined by structural studies on the mutant W168F and its complexes with ligands. The three-dimensional structures of unliganded mutant (1.8 A) and complexes with sulfate (2.8 A) and glycerol-2-phosphate (G2P) (2.8 A) have been determined. Loop6 was found disordered in these structures, reflecting the importance of W168 in stabilizing either the open or the closed states. Critical sequence differences between the Plasmodium enzyme and other TIMs may influence the equilibrium between the closed and open forms. Examination of the environment of the loop6 shows that its propensity for the open or the closed forms is influenced not only by Phe96 as suggested earlier, but also by Asn233, which occurs in the vicinity of the active site. This residue is Gly in the other TIM sequences and probably plays a crucial role in the mode of ligand binding, which in turn affects the loop opening/closing process in PfTIM.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: