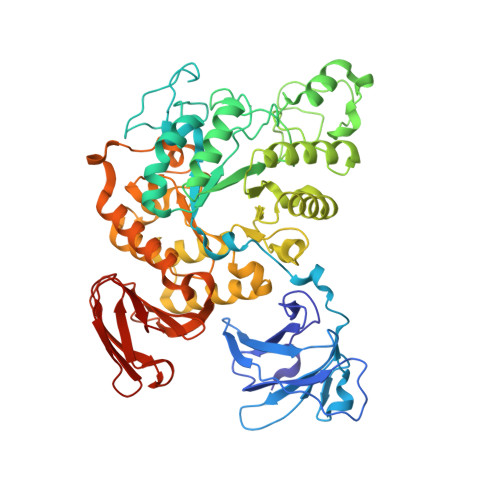
Complex structures of Thermoactinomyces vulgaris R-47 alpha-amylase 2 with acarbose and cyclodextrins demonstrate the multiple substrate recognition mechanism
Ohtaki, A., Mizuno, M., Tonozuka, T., Sakano, Y., Kamitori, S.(2004) J Biological Chem 279: 31033-31040
- PubMed: 15138257 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M404311200
- Primary Citation Related Structures:
1VFM, 1VFO, 1VFU, 3A6O - PubMed Abstract:
Thermoactinomyces vulgaris R-47 alpha-amylase 2 (TVAII) has the unique ability to hydrolyze cyclodextrins (CDs), with various sized cavities, as well as starch. To understand the relationship between structure and substrate specificity, x-ray structures of a TVAII-acarbose complex and inactive mutant TVAII (D325N/D421N)/alpha-, beta- and gamma-CDs complexes were determined at resolutions of 2.9, 2.9, 2.8, and 3.1 A, respectively. In all complexes, the interactions between ligands and enzymes at subsites -1, -2, and -3 were almost the same, but striking differences in the catalytic site structure were found at subsites +1 and +2, where Trp(356) and Tyr(374) changed the conformation of the side chain depending on the structure and size of the ligands. Trp(356) and Tyr(374) are thought to be responsible for the multiple substrate-recognition mechanism of TVAII, providing the unique substrate specificity. In the beta-CD complex, the beta-CD maintains a regular conical structure, making it difficult for Glu(354) to protonate the O-4 atom at the hydrolyzing site as a previously proposed hydrolyzing mechanism of alpha-amylase. From the x-ray structures, it is suggested that the protonation of the O-4 atom is possibly carried out via a hydrogen atom of the inter-glucose hydrogen bond at the hydrolyzing site.
- Department of Biotechnology and Life Science, Tokyo University of Agriculture and Technology, 2-24-16 Naka-cho, Koganei, Tokyo 184-8588, Japan.
Organizational Affiliation: