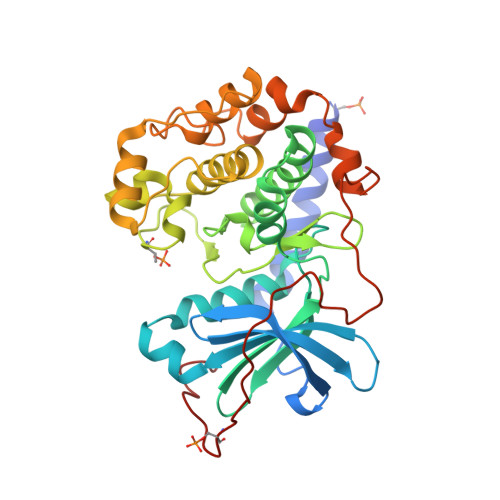
Structure-based optimization of novel azepane derivatives as PKB inhibitors
Breitenlechner, C.B., Wegge, T., Berillon, L., Graul, K., Marzenell, K., Friebe, W.-G., Thomas, U., Schumacher, R., Huber, R., Engh, R.A., Masjost, B.(2004) J Med Chem 47: 1375-1390
- PubMed: 14998327 Search on PubMed
- DOI: https://doi.org/10.1021/jm0310479
- Primary Citation Related Structures:
1SVE, 1SVG, 1SVH, 1VEB - PubMed Abstract:
Novel azepane derivatives were prepared and evaluated for protein kinase B (PKB-alpha) and protein kinase A (PKA) inhibition. The original (-)-balanol-derived lead structure (4R)-4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-benzoic acid (3R)-3-[(pyridine-4-carbonyl)amino]-azepan-4-yl ester (1) (IC(50) (PKB-alpha) = 5 nM) which contains an ester moiety was found to be plasma unstable and therefore unsuitable as a drug. Based upon molecular modeling studies using the crystal structure of the complex between PKA and 1, the five compounds N-[(3R,4R)-4-[4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-benzoylamino]-azepan-3-yl]-isonicotinamide (4), (3R,4R)-N-[4-[4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-benzyloxy]-azepan-3-yl]-isonicotinamide (5), N-[(3R,4S)-4-[4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-phenylamino]-methyl]-azepan-3-yl)-isonicotinamide (6), N-[(3R,4R)-4-[4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-benzylamino]-azepan-3-yl]-isonicotinamide (7), and N-[(3R,4S)-4-(4-[trans-2-[4-(2-fluoro-6-hydroxy-3-methoxy-benzoyl)-phenyl]-vinyl]-azepan-3-yl)-isonicotinamide (8) with linkers isosteric to the ester were designed, synthesized, and tested for in vitro inhibitory activity against PKA and PKB-alpha and for plasma stability in mouse plasma.(1) Compound 4 was found to be plasma stable and highly active (IC(50) (PKB-alpha) = 4 nM). Cocrystals with PKA were obtained for 4, 5, and 8 and analyzed for binding interactions and conformational changes in the ligands and protein in order to rationalize the different activities of the molecules.
- Pharma Research, Roche Diagnostics GmbH, Werk Penzberg, Nonnenwald 2, D-82372 Penzberg, Germany. birgit.masjost@roche.com
Organizational Affiliation: