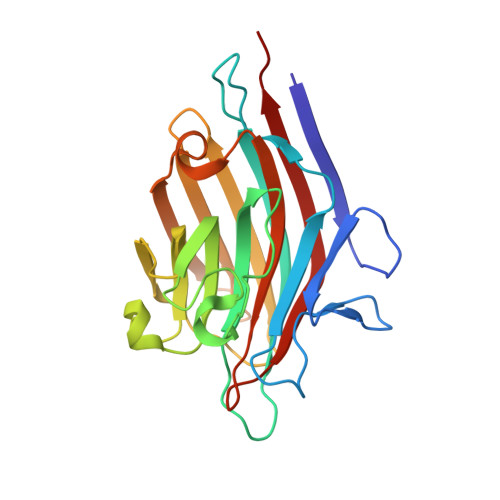
Structural plasticity of peanut lectin: an X-ray analysis involving variation in pH, ligand binding and crystal structure.
Kundhavai Natchiar, S., Arockia Jeyaprakash, A., Ramya, T.N., Thomas, C.J., Suguna, K., Surolia, A., Vijayan, M.(2004) Acta Crystallogr D Biol Crystallogr 60: 211-219
- PubMed: 14747696 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490302849X
- Primary Citation Related Structures:
1V6I, 1V6J, 1V6K, 1V6L, 1V6M, 1V6N, 1V6O - PubMed Abstract:
Until recently, it has only been possible to grow crystals of peanut lectin when complexed with sugar ligands. It is now shown that it is possible to grow peanut lectin crystals at acidic pH in the presence of oligopeptides corresponding to a loop in the lectin molecule. Crystals have also been prepared in the presence of these peptides as well as lactose. Low-pH crystal forms of the lectin-lactose complex similar to those obtained at neutral pH have also been grown. Thus, crystals of peanut lectin grown under different environmental conditions, at two pH values with and without sugar bound to the lectin, are now available. They have been used to explore the plasticity and hydration of the molecule. A detailed comparison between different structures shows that the lectin molecule is sturdy and that the effect of changes in pH, ligand binding and environment on it is small. The region involving the curved front beta-sheet and the loops around the second hydrophobic core is comparatively rigid. The back beta-sheet involved in quaternary association, which exhibits considerable variability, is substantially flexible, as is the sugar-binding region. The numbers of invariant water molecules in the hydration shell are small and they are mainly involved in metal coordination or in stabilizing unusual structural features. Small consistent movements occur in the combining site upon sugar binding, although the site is essentially preformed.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: