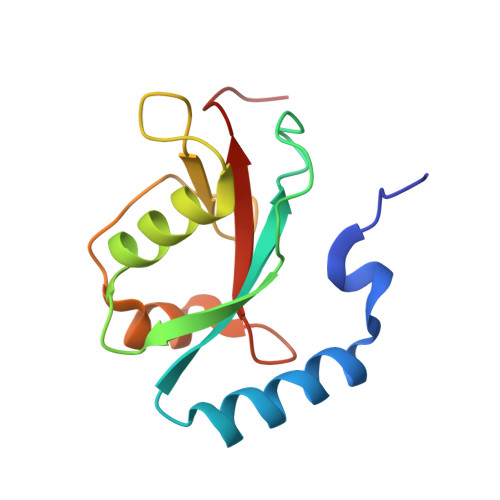
Solution structure of microtubule-associated protein light chain 3 and identification of its functional subdomains.
Kouno, T., Mizuguchi, M., Tanida, I., Ueno, T., Kanematsu, T., Mori, Y., Shinoda, H., Hirata, M., Kominami, E., Kawano, K.(2005) J Biological Chem 280: 24610-24617
- PubMed: 15857831 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M413565200
- Primary Citation Related Structures:
1V49 - PubMed Abstract:
Microtubule-associated protein (MAP) light chain 3 (LC3) is a human homologue of yeast Apg8/Aut7/Cvt5 (Atg8), which is essential for autophagy. MAP-LC3 is cleaved by a cysteine protease to produce LC3-I, which is located in cytosolic fraction. LC3-I, in turn, is converted to LC3-II through the actions of E1- and E2-like enzymes. LC3-II is covalently attached to phosphatidylethanolamine on its C terminus, and it binds tightly to autophagosome membranes. We determined the solution structure of LC3-I and found that it is divided into N- and C-terminal subdomains. Additional analysis using a photochemically induced dynamic nuclear polarization technique also showed that the N-terminal subdomain of LC3-I makes contact with the surface of the C-terminal subdomain and that LC3-I adopts a single compact conformation in solution. Moreover, the addition of dodecylphosphocholine into the LC3-I solution induced chemical shift perturbations primarily in the C-terminal subdomain, which implies that the two subdomains have different sensitivities to dodecylphosphocholine micelles. On the other hand, deletion of the N-terminal subdomain abolished binding of tubulin and microtubules. Thus, we showed that two subdomains of the LC3-I structure have distinct functions, suggesting that MAP-LC3 can act as an adaptor protein between microtubules and autophagosomes.
- Faculty of Pharmaceutical Sciences, Toyama Medical and Pharmaceutical University, Toyama 930-0194, Japan.
Organizational Affiliation: