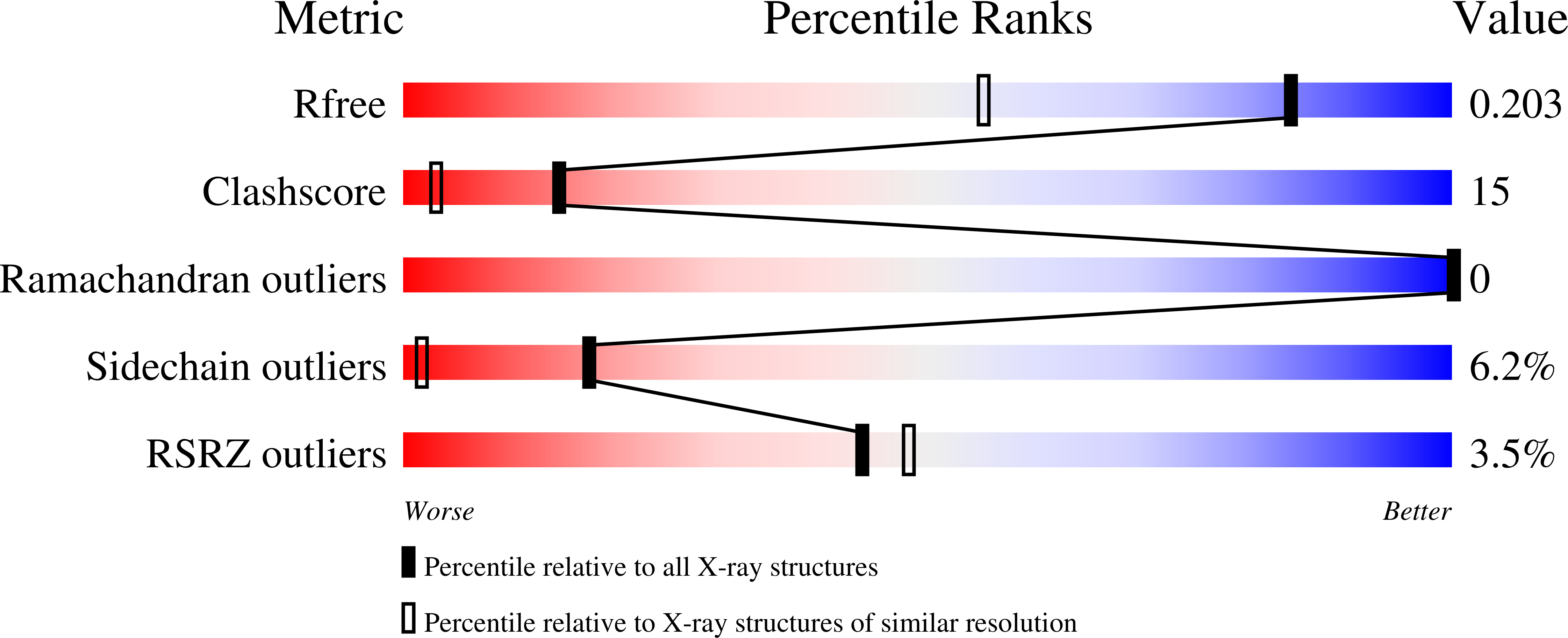
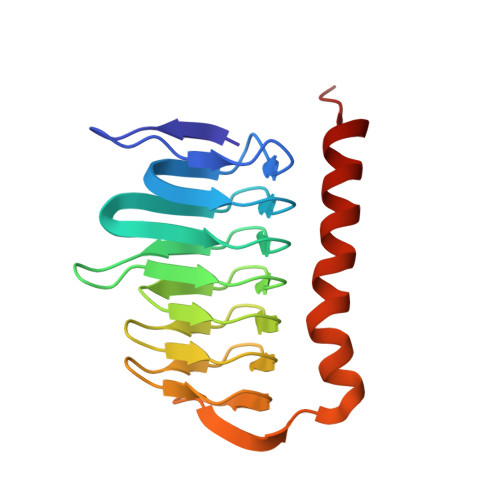
Observation of a calcium-binding site in the gamma-class carbonic anhydrase from Pyrococcus horikoshii.
Jeyakanthan, J., Rangarajan, S., Mridula, P., Kanaujia, S.P., Shiro, Y., Kuramitsu, S., Yokoyama, S., Sekar, K.(2008) Acta Crystallogr D Biol Crystallogr 64: 1012-1019
- PubMed: 18931408 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908024323
- Primary Citation Related Structures:
1V3W, 1V67, 2FKO - PubMed Abstract:
Carbonic anhydrases are zinc-containing metalloenzymes that catalyze the interconversion of carbon dioxide and bicarbonate. Three crystal structures of gamma-class carbonic anhydrase (one of which is bound to a bicarbonate molecule) from the aerobic OT3 strain of the hyperthermophilic archeon Pyrococcus horikoshii have been solved by molecular replacement in space group F4(1)32. The asymmetric unit contains a monomer of 173 amino acids and a catalytic Zn2+ ion. The protein fold is a regular prism formed by a left-handed beta-helix, similar to previously reported structures. The active-site Zn2+ ion located at the interface between the two monomers is bound to three histidyl residues and a water molecule in a tetrahedral fashion. In addition to the 20 beta-strands comprising the beta-helix, there is also a long C-terminal alpha-helix. For the first time, Ca2+ ions have been observed in addition to the catalytic Zn2+ ion. It is hypothesized that Tyr159 (which corresponds to the catalytically important Asn202 in previously reported structures) utilizes C-H...pi interactions to fulfill its functions. This study may shed light on the catalytic mechanism of the enzyme and throw open new questions on the mechanism of product removal in carbonic anhydrases.
- National Synchrotron Radiation Research Center, 101 Hsin-Ann Road, Hsinchu Science Park, Hsinchu 30076, Taiwan.
Organizational Affiliation: