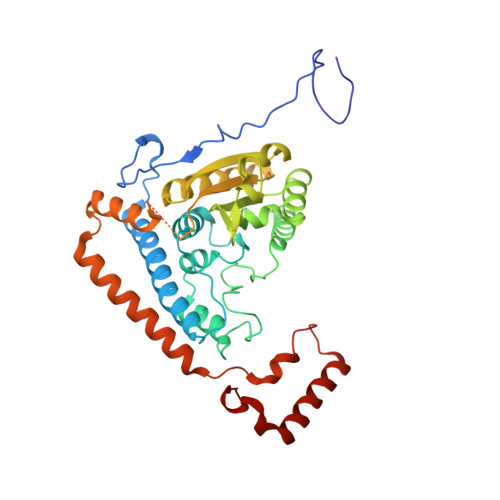
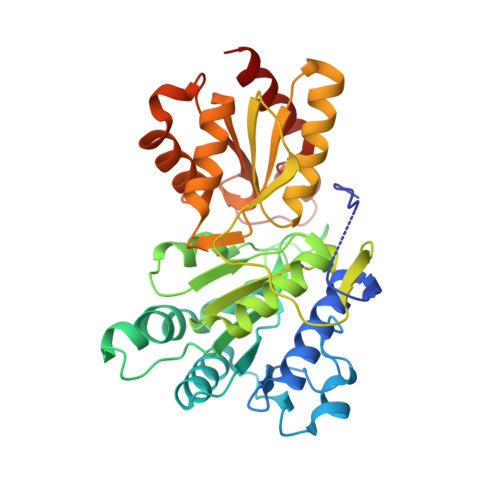
Cross-Talk between Thiamin Diphosphate Binding and Phosphorylation Loop Conformation in Human Branched-Chain {Alpha}-Keto Acid Decarboxylase/Dehydrogenase
Li, J., Wynn, R.M., Machius, M., Chuang, J.L., Karthikeyan, S., Tomchick, D.R., Chuang, D.T.(2004) J Biological Chem 279: 32968
- PubMed: 15166214 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M403611200
- Primary Citation Related Structures:
1V11, 1V16, 1V1M, 1V1R - PubMed Abstract:
The decarboxylase/dehydrogenase (E1b) component of the 4-megadalton human branched-chain alpha-keto acid dehydrogenase (BCKD) metabolic machine is a thiamin diphosphate (ThDP)-dependent enzyme with a heterotetrameric cofactor-binding fold. The E1b component catalyzes the decarboxylation of alpha-keto acids and the subsequent reductive acylation of the lipoic acid-bearing domain (LBD) from the 24-meric transacylase (E2b) core. In the present study, we show that the binding of cofactor ThDP to the E1b active site induces a disorder-to-order transition of the conserved phosphorylation loop carrying the two phosphorylation sites Ser(292)-alpha and Ser(302)-alpha, as deduced from the 1.80-1.85 A apoE1b and holoE1b structures. The induced loop conformation is essential for the recognition of lipoylated LBD to initiate E1b-catalyzed reductive acylation. Alterations of invariant Arg(287)-alpha, Asp(295)-alpha, Tyr(300)-alpha, and Arg(301)-alpha that form a hydrogen-bonding network in the phosphorylation loop result in the disordering of the loop conformation as elucidated by limited proteolysis, accompanied by the impaired binding and diminished reductive acylation of lipoylated LBD. In contrast, k(cat) values for E1b-catalyzed decarboxylation of the alpha-keto acid are higher in these E1b mutants than in wild-type E1b, with higher K(m) values for the substrate in the mutants. ThDP binding that orders the loop prevents phosphorylation of E1b by the BCKD kinase and averts the inactivation of wild-type E1b, but not the above mutants, by this covalent modification. Our results establish that the cross-talk between the bound ThDP and the phosphorylation loop conformation serves as a feed-forward switch for multiple reaction steps in the BCKD metabolic machine.
- Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9038, USA.
Organizational Affiliation: