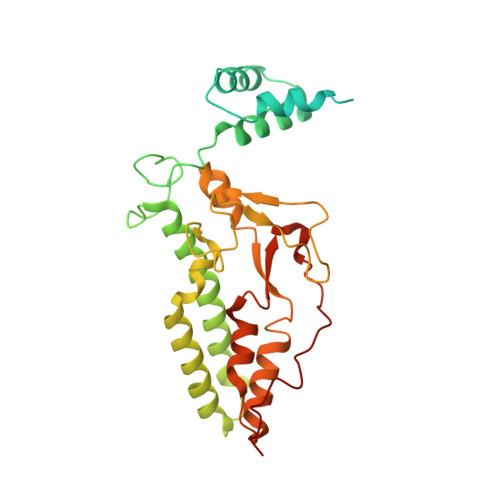
Structural Mechanism for Inactivation and Activation of Cad/Dff40 in the Apoptotic Pathway
Woo, E.-J., Kim, Y.-G., Kim, M.-S., Han, W.-D., Shin, S., Robinson, H., Park, S.-Y., Oh, B.-H.(2004) Mol Cell 14: 531
- PubMed: 15149602 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(04)00258-8
- Primary Citation Related Structures:
1V0D - PubMed Abstract:
CAD/DFF40 is responsible for the degradation of chromosomal DNA into nucleosomal fragments and subsequent chromatin condensation during apoptosis. It exists as an inactive complex with its inhibitor ICAD/DFF45 in proliferating cells but becomes activated upon cleavage of ICAD/DFF45 into three domains by caspases in dying cells. The molecular mechanism underlying the control and activation of CAD/DFF40 was unknown. Here, the crystal structure of activated CAD/DFF40 reveals that it is a pair of molecular scissors with a deep active-site crevice that appears ideal for distinguishing internucleosomal DNA from nucleosomal DNA. Ensuing studies show that ICAD/DFF45 sequesters the nonfunctional CAD/DFF40 monomer and is also able to disassemble the functional CAD/DFF40 dimer. This capacity requires the involvement of the middle domain of ICAD/DFF45, which by itself cannot remain bound to CAD/DFF40 due to low binding affinity for the enzyme. Thus, the consequence of the caspase-cleavage of ICAD/DFF45 is a self-assembly of CAD/DFF40 into the active dimer.
- Division of Molecular and Life Sciences, Department of Life Sciences and Center for Biomolecular Recognition, Pohang University of Science and Technology, Pohang, Kyungbuk 790-784, South Korea.
Organizational Affiliation: