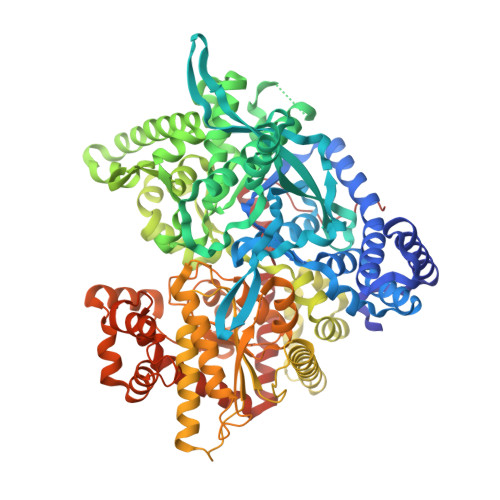
Binding of the potential antitumour agent indirubin-5-sulphonate at the inhibitor site of rabbit muscle glycogen phosphorylase b. Comparison with ligand binding to pCDK2-cyclin A complex.
Kosmopoulou, M.N., Leonidas, D.D., Chrysina, E.D., Bischler, N., Eisenbrand, G., Sakarellos, C.E., Pauptit, R., Oikonomakos, N.G.(2004) Eur J Biochem 271: 2280-2290
- PubMed: 15153119 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.2004.04173.x
- Primary Citation Related Structures:
1UZU - PubMed Abstract:
The binding of indirubin-5-sulphonate (E226), a potential anti-tumour agent and a potent inhibitor (IC(50) = 35 nm) of cyclin-dependent kinase 2 (CDK2) and glycogen phosphorylase (GP) has been studied by kinetic and crystallographic methods. Kinetic analysis revealed that E226 is a moderate inhibitor of GPb (K(i) = 13.8 +/- 0.2 micro m) and GPa (K(i) = 57.8 +/- 7.1 micro m) and acts synergistically with glucose. To explore the molecular basis of E226 binding we have determined the crystal structure of the GPb/E226 complex at 2.3 A resolution. Structure analysis shows clearly that E226 binds at the purine inhibitor site, where caffeine and flavopiridol also bind [Oikonomakos, N.G., Schnier, J.B., Zographos, S.E., Skamnaki, V.T., Tsitsanou, K.E. & Johnson, L.N. (2000) J. Biol. Chem.275, 34566-34573], by intercalating between the two aromatic rings of Phe285 and Tyr613. The mode of binding of E226 to GPb is similar, but not identical, to that of caffeine and flavopiridol. Comparative structural analyses of the GPb-E226, GPb-caffeine and GPb-flavopiridol complex structures reveal the structural basis of the differences in the potencies of the three inhibitors and indicate binding residues in the inhibitor site that can be exploited to obtain more potent inhibitors. Structural comparison of the GPb-E226 complex structure with the active pCDK2-cyclin A-E226 complex structure clearly shows the different binding modes of the ligand to GPb and CDK2; the more extensive interactions of E226 with the active site of CDK2 may explain its higher affinity towards the latter enzyme.
- Institute of Organic and Pharmaceutical Chemistry, The National Hellenic Research Foundation, 48 Vas. Constantinou Avenue, 11635 Athens, Greece.
Organizational Affiliation: