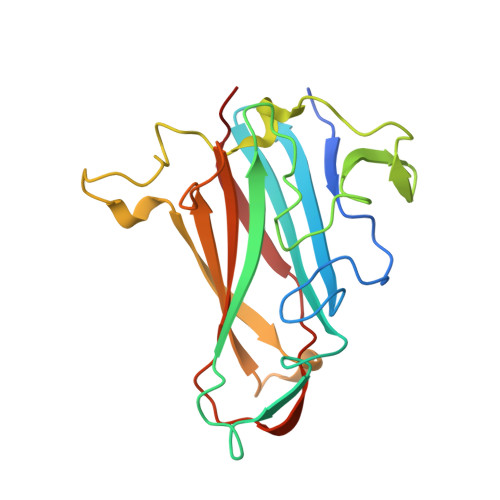
Crystal Structure of Species D Adenovirus Fiber Knobs and Their Sialic Acid Binding Sites
Burmeister, W.P., Guilligay, D., Cusack, S., Wadell, G., Arnberg, N.(2004) J Virol 78: 7727
- PubMed: 15220447 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.78.14.7727-7736.2004
- Primary Citation Related Structures:
1UXA, 1UXB, 1UXE - PubMed Abstract:
Adenovirus serotype 37 (Ad37) belongs to species D and can cause epidemic keratoconjunctivitis, whereas the closely related Ad19p does not. Primary cell attachment by adenoviruses is mediated through receptor binding of the knob domain of the fiber protein. The knobs of Ad37 and Ad19p differ at only two positions, Lys240Glu and Asn340Asp. We report the high-resolution crystal structures of the Ad37 and Ad19p knobs, both native and in complex with sialic acid, which has been proposed as a receptor for Ad37. Overall, the Ad37 and Ad19p knobs are very similar to previously reported knob structures, especially to that of Ad5, which binds the coxsackievirus-adenovirus receptor (CAR). Ad37 and Ad19p knobs are structurally identical with the exception of the changed side chains and are structurally most similar to CAR-binding knobs (e.g., that of Ad5) rather than non-CAR-binding knobs (e.g., that of Ad3). The two mutations in Ad19p result in a partial loss of the exceptionally high positive surface charge of the Ad37 knob but do not affect sialic acid binding. This site is located on the top of the trimer and binds both alpha(2,3) and alpha(2,6)-linked sialyl-lactose, although only the sialic acid residue makes direct contact. Amino acid alignment suggests that the sialic acid binding site is conserved in several species D serotypes. Our results show that the altered viral tropism and cell binding of Ad19p relative to those of Ad37 are not explained by a different binding ability toward sialyl-lactose.
- Laboratoire de Virologie Moléculaire et Structurale, Université Joseph Fourier, Faculté de Médecine de Grenoble, F-38042 Grenoble cedex 9, France.
Organizational Affiliation: