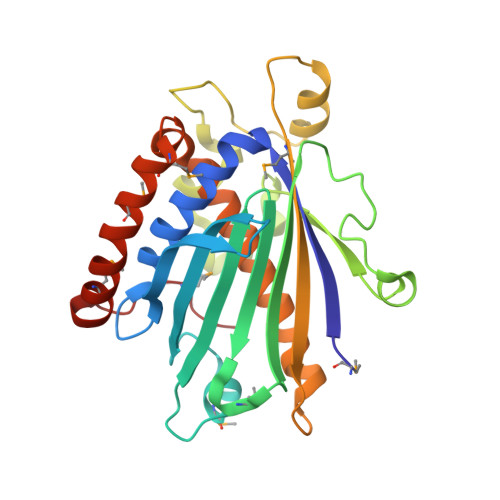
Structure-Function Analysis of Human [Corrected] Phosphatidylinositol Transfer Protein Alpha Bound to Phosphatidylinositol.
Tilley, S.J., Skippen, A., Murray-Rust, J., Swigart, P.M., Stewart, A., Morgan, C.P., Cockcroft, S., Mcdonald, N.Q.(2004) Structure 12: 317
- PubMed: 14962392 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.01.013
- Primary Citation Related Structures:
1UW5 - PubMed Abstract:
Phosphatidylinositol transfer protein alpha (PITPalpha) selectively transports and promotes exchange of phosphatidylinositol (PI) and phosphatidylcholine (PC) between lipid bilayers. In higher eukaryotes PITPalpha is required for cellular functions such as phospholipase C-mediated signaling, regulated exocytosis, and secretory vesicle formation. We have determined the crystal structure of human PITPalpha bound to its physiological ligand, PI, at 2.95 A resolution. The structure identifies the critical side chains within the lipid-headgroup binding pocket that define the exquisite specificity for PI. Mutational analysis of the PI binding pocket is in good agreement with the structural data and allows manipulation of functional properties of PITPalpha. Surprisingly, there are no major conformational differences between PI- and PC-loaded PITPalpha, despite previous predictions. In the crystal, PITPalpha-PI is dimeric, with two identical dimers in the asymmetric unit. The dimer interface masks precisely the sequence we identify as contributing to PITPalpha membrane interaction. Our structure represents a soluble, transport-competent form of PI-loaded PITPalpha.
- School of Crystallography, Birkbeck, University of London, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: