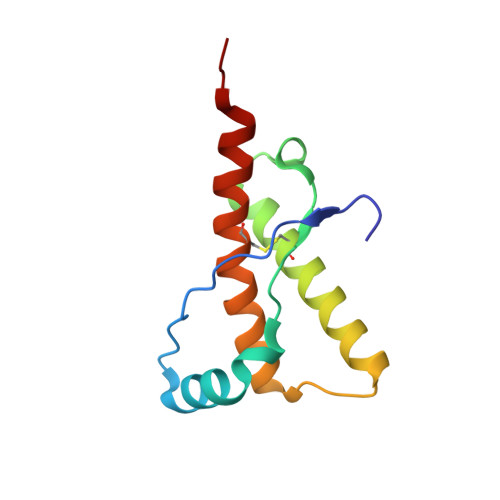
The Crystal Structure of the Globular Domain of Sheep Prion Protein
Haire, L.F., Whyte, S.M., Vasisht, N., Gill, A.C., Verma, C., Dodson, E.J., Dodson, G.G., Bayley, P.M.(2004) J Mol Biology 336: 1175
- PubMed: 15037077 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.12.059
- Primary Citation Related Structures:
1UW3 - PubMed Abstract:
The prion protein PrP is a naturally occurring polypeptide that becomes transformed from a normal conformation to that of an aggregated form, characteristic of pathological states in fatal transmissible spongiform conditions such as Creutzfeld-Jacob Disease and Bovine Spongiform Encephalopathy. We report the crystal structure, at 2 A resolution, of residues 123-230 of the C-terminal globular domain of the ARQ allele of sheep prion protein (PrP). The asymmetric unit contains a single molecule whose secondary structure and overall organisation correspond to those structures of PrPs from various mammalian species determined by NMR. The globular domain shows a close association of helix-1, the C-terminal portion of helix-2 and the N-terminal portion of helix-3, bounded by the intramolecular disulphide bond, 179-214. The loop 164-177, between beta2 and helix-2 is relatively well structured compared to the human PrP NMR structure. Analysis of the sheep PrP structure identifies two possible loci for the initiation of beta-sheet mediated polymerisation. One of these comprises the beta-strand, residues 129-131 that forms an intra-molecular beta-sheet with residues 161-163. This strand is involved in lattice contacts about a crystal dyad to generate a four-stranded intermolecular beta-sheet between neighbouring molecules. The second locus involves the region 188-204, which modelling suggests is able to undergo a partial alpha-->beta switch within the monomer. These loci provide sites within the PrPc monomer that could readily give rise to early intermediate species on the pathway to the formation of aggregated PrPSc containing additional intermolecular beta-structure.
- Structural Biology Group, Division of Protein Structure, National Institute for Medical Research, The Ridgeway, Mill Hill, London NW7 1AA, UK.
Organizational Affiliation: