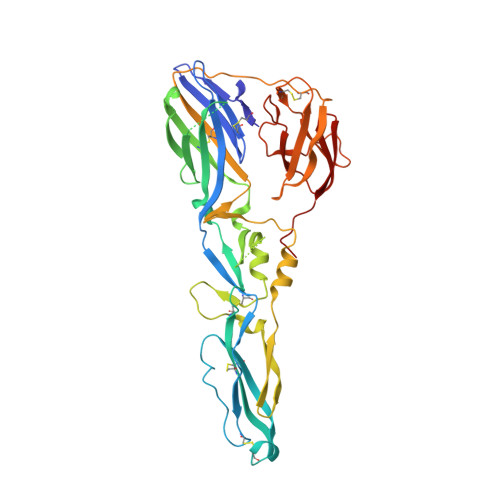
Structure of a Flavivirus Envelope Glycoprotein in its Low-Ph-Induced Membrane Fusion Conformation.
Bressanelli, S., Stiasny, K., Allison, S.L., Stura, E.A., Duquerroy, S., Lescar, J., Heinz, F.X., Rey, F.A.(2004) EMBO J 23: 728
- PubMed: 14963486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600064
- Primary Citation Related Structures:
1URZ - PubMed Abstract:
Enveloped viruses enter cells via a membrane fusion reaction driven by conformational changes of specific viral envelope proteins. We report here the structure of the ectodomain of the tick-borne encephalitis virus envelope glycoprotein, E, a prototypical class II fusion protein, in its trimeric low-pH-induced conformation. We show that, in the conformational transition, the three domains of the neutral-pH form are maintained but their relative orientation is altered. Similar to the postfusion class I proteins, the subunits rearrange such that the fusion peptide loops cluster at one end of an elongated molecule and the C-terminal segments, connecting to the viral transmembrane region, run along the sides of the trimer pointing toward the fusion peptide loops. Comparison with the low-pH-induced form of the alphavirus class II fusion protein reveals striking differences at the end of the molecule bearing the fusion peptides, suggesting an important conformational effect of the missing membrane connecting segment.
- Virologie Moléculaire & Structurale, CNRS UMR 2472/INRA UMR 1157, IFR 115 Gif-sur-Yvette, France.
Organizational Affiliation: