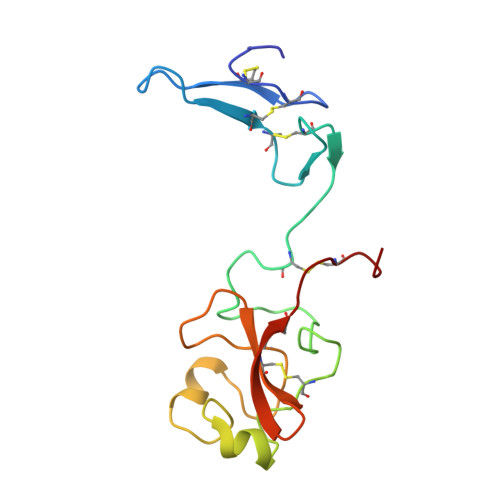
Solution structure of the amino-terminal fragment of urokinase-type plasminogen activator.
Hansen, A.P., Petros, A.M., Meadows, R.P., Nettesheim, D.G., Mazar, A.P., Olejniczak, E.T., Xu, R.X., Pederson, T.M., Henkin, J., Fesik, S.W.(1994) Biochemistry 33: 4847-4864
- PubMed: 8161544 Search on PubMed
- DOI: https://doi.org/10.1021/bi00182a013
- Primary Citation Related Structures:
1URK - PubMed Abstract:
The amino-terminal fragment (ATF) of urokinase-type plasminogen activator is a two domain protein which consists of a growth factor and a kringle domain. The 1H, 13C, and 15N chemical shifts of this protein have been assigned using heteronuclear two- and three-dimensional NMR experiments on selective and uniformly 15N- and 15N/13C-labeled protein isolated from mammalian cells that overexpress the protein. The chemical shift assignments were used to interpret the NOE data which resulted in a total of 1299 NOE restraints. The NOE restraints were used along with 27 phi angle restraints and 21 hydrogen-bonding restraints to produce 15 low energy structures. The individual domains in the structures are highly converged, but the two domains are structurally independent. The root mean square deviations (rmsd) between residues 11-46 in the growth factor domain and the mean atomic coordinates were 0.99 +/- 0.2 for backbone heavy atoms and 1.65 +/- 0.2 for all non-hydrogen atoms. For residues 55-130 in the kringle domain, the rmsd was 0.84 +/- 0.2 for backbone heavy atoms and 1.42 +/- 0.2 for all non-hydrogen atoms. The overall structures of the individual domains are very similar to the structures of homologous proteins. However, important structural differences between the growth factor and other homologous proteins were observed in the region which has been implicated in binding the urokinase receptor which may explain, in part, why other growth factors show no appreciable affinity for the urokinase receptor.
- Pharmaceutical Discovery Division, Abbott Laboratories, Abbott Park, Illinois 60064.
Organizational Affiliation: