Molecular Dissection of the Interaction between the Small G Proteins Rac1 and Rhoa and Protein Kinase C-Related Kinase 1 (Prk1)
Owen, D., Lowe, P.N., Nietlispach, D., Brosnan, C.E., Chirgadze, D.Y., Parker, P.J., Blundell, T.L., Mott, H.R.(2003) J Biological Chem 278: 50578
- PubMed: 14514689 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304313200
- Primary Citation Related Structures:
1URF - PubMed Abstract:
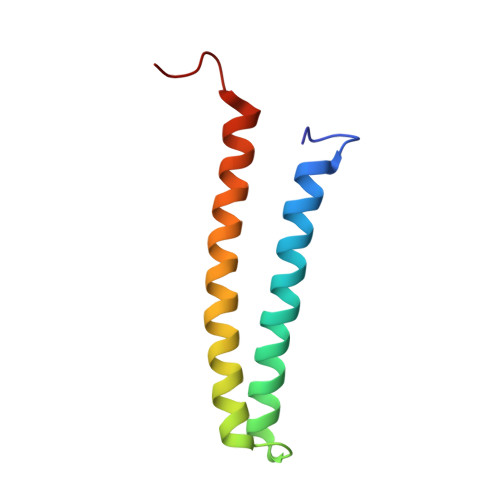
PRK1 is a serine/threonine kinase that belongs to the protein kinase C superfamily. It can be activated either by members of the Rho family of small G proteins, by proteolysis, or by interaction with lipids. Here we investigate the binding of PRK1 to RhoA and Rac1, two members of the Rho family. We demonstrate that PRK1 binds with a similar affinity to RhoA and Rac1. We present the solution structure of the second HR1 domain from the regulatory N-terminal region of PRK1, and we show that it forms an anti-parallel coiled-coil. In addition, we have used NMR to map the binding contacts of the HR1b domain with Rac1. These are compared with the contacts known to form between HR1a and RhoA. We have used mutagenesis to define the residues in Rac that are important for binding to HR1b. Surprisingly, as well as residues adjacent to Switch I, in Switch II, and in helix alpha5, it appears that the C-terminal stretch of basic amino acids in Rac is required for a high affinity interaction with HR1b.
- Department of Biochemistry, University of Cambridge, Tennis Court Road, Cambridge CB2 1GA, United Kingdom. do@bioc.cam.ac.uk
Organizational Affiliation: