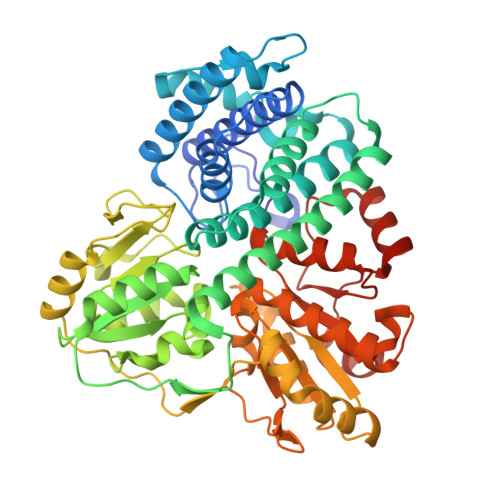
Structure of the Hybrid Cluster Protein (Hcp) from Desulfovibrio Desulfuricans Atcc 27774 Containing Molecules in the Oxidized and Reduced States
Macedo, S., Aragao, D., Mitchell, E.P., Lindley, P.F.(2003) Acta Crystallogr D Biol Crystallogr 59: 2065
- PubMed: 14646063 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903025861
- Primary Citation Related Structures:
1UPX - PubMed Abstract:
The hybrid cluster protein (HCP) from the sulfate-reducing bacteria Desulfovibrio desulfuricans ATCC 27774 has been isolated and crystallized anaerobically. The protein sample used in the crystallization studies was several months old, having been stored at 193 K, and initial crystal structure studies were unable to fully resolve details of the hybrid cluster despite the use of high-resolution data to 1.25 A collected at the ESRF, Grenoble, France. Full elucidation of the structure has only become possible with the complete knowledge of the as-isolated and fully reduced crystal structures. The analysis clarifies the significant movements in the position of the Fe atom linked to the persulfide moiety in the oxidized as-isolated protein and the S atom of the persulfide itself as the protein is reduced. The structures of the as-isolated and reduced states are discussed in terms of the putative function of the HCP proteins.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Av. República, EAN, Apartado 127, 2781-901 Oeiras, Portugal.
Organizational Affiliation: