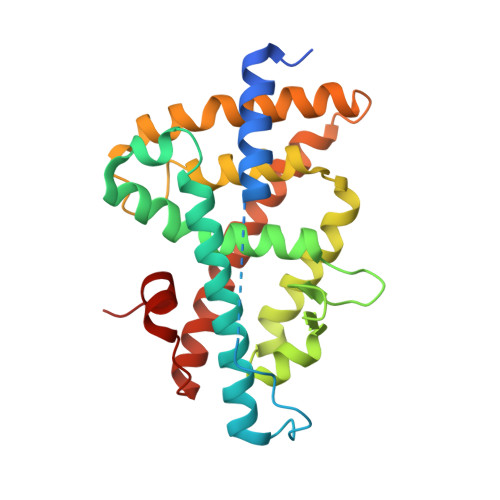
Crystal Structure of the Human Liver X Receptor Beta Ligand-Binding Domain in Complex with a Synthetic Agonist
Hoerer, S., Schmid, A., Heckel, A., Budzinski, R.M., Nar, H.(2003) J Mol Biology 334: 853
- PubMed: 14643652 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.033
- Primary Citation Related Structures:
1UPV, 1UPW - PubMed Abstract:
LXRbeta belongs to the nuclear hormone receptor superfamily of ligand-activated transcription factors. Its natural ligands are supposed to be oxidised derivatives of cholesterol. Stimulation of LXRbeta by agonists activates a number of genes that are involved in the regulation of lipid metabolism and cholesterol efflux from cells. Therefore, LXRbeta may represent a novel therapeutic target for the treatment of dyslipidemia and atherosclerosis.Here, we report the X-ray crystal structure of the LXRbeta ligand-binding domain in complex with a synthetic agonist, T-0901317. This compound occupies the ligand-binding pocket of the receptor, forms numerous lipophilic contacts with the protein and one crucial hydrogen bond to His435 and stabilises the agonist conformation of the receptor ligand-binding domain. The recruitment of the AF2-region of the protein is not achieved via direct polar interactions of the ligand with protein side-chains of this helical segment, but rather via few hydrophobic contacts and probably more importantly via indirect effects involving the pre-orientation of side-chains that surround the ligand-binding pocket and form the interface to the AF2-helix. On the basis of these results we propose a binding mode and a mechanism of action for the putative natural ligands, oxidised derivatives of cholesterol.
- Department of Lead Discovery, Boehringer Ingelheim Pharma GmbH & Co. KG, Birkendorfer Strasse 65, D-88397 Biberach/Riss, Germany.
Organizational Affiliation: