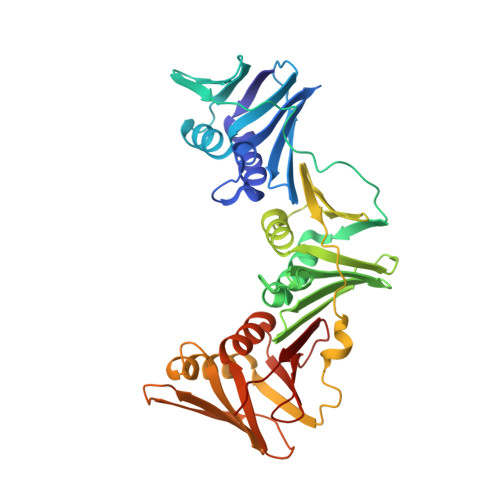
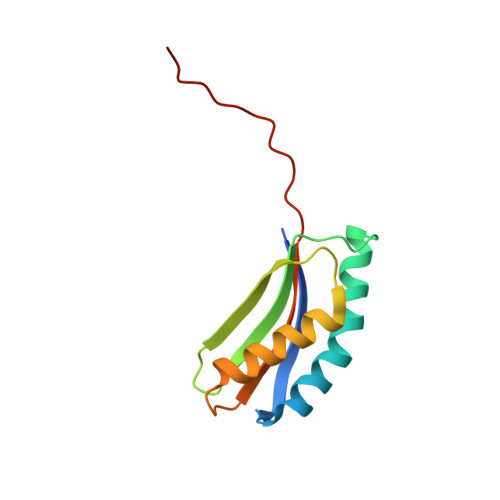
Structural Basis for Recruitment of Translesion DNA Polymerase Pol Iv/Dinb to the Beta-Clamp
Bunting, K.A., Roe, S.M., Pearl, L.H.(2003) EMBO J 22: 5883
- PubMed: 14592985
- DOI: https://doi.org/10.1093/emboj/cdg568
- Primary Citation Related Structures:
1UNN - PubMed Abstract:
Y-family DNA polymerases can extend primer strands across template strand lesions that stall replicative polymerases. The poor processivity and fidelity of these enzymes, key to their biological role, requires that their access to the primer-template junction is both facilitated and regulated in order to minimize mutations. These features are believed to be provided by interaction with processivity factors, beta-clamp or proliferating cell nuclear antigen (PCNA), which are also essential for the function of replicative DNA polymerases. The basis for this interaction is revealed by the crystal structure of the complex between the 'little finger' domain of the Y-family DNA polymerase Pol IV and the beta-clamp processivity factor, both from Escherichia coli. The main interaction involves a C-terminal peptide of Pol IV, and is similar to interactions seen between isolated peptides and other processivity factors. However, this first structure of an entire domain of a binding partner with an assembled clamp reveals a substantial secondary interface, which maintains the polymerase in an inactive orientation, and may regulate the switch between replicative and Y-family DNA polymerases in response to a template strand lesion.
- The Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: