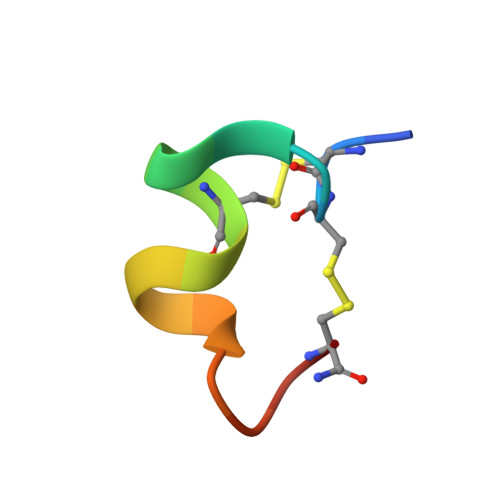
Solution conformation of alpha-conotoxin GIC, a novel potent antagonist of alpha3beta2 nicotinic acetylcholine receptors
Chi, S.-W., Kim, D.-H., Olivera, B.M., McIntosh, J.M., Han, K.-H.(2004) Biochem J 380: 347-352
- PubMed: 14992691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20031792
- Primary Citation Related Structures:
1UL2 - PubMed Abstract:
Alpha-conotoxin GIC is a 16-residue peptide isolated from the venom of the cone snail Conus geographus. Alpha-conotoxin GIC potently blocks the alpha3beta2 subtype of human nicotinic acetylcholine receptor, showing a high selectivity for neuronal versus muscle subtype [McIntosh, Dowell, Watkins, Garrett, Yoshikami, and Olivera (2002) J. Biol. Chem. 277, 33610-33615]. We have now determined the three-dimensional solution structure of alpha-conotoxin GIC by NMR spectroscopy. The structure of alpha-conotoxin GIC is well defined with backbone and heavy atom root mean square deviations (residues 2-16) of 0.53 A and 0.96 A respectively. Structure and surface comparison of alpha-conotoxin GIC with the other alpha4/7 subfamily conotoxins reveals unique structural aspects of alpha-conotoxin GIC. In particular, the structural comparison between alpha-conotoxins GIC and MII indicates molecular features that may confer their similar receptor specificity profile, as well as those that provide the unique binding characteristics of alpha-conotoxin GIC.
- Laboratory of Protein Analysis and Design, Division of Drug Discovery, Korea Research Institute of Bioscience and Biotechnology, Yusong P.O. Box 115, Daejon, Korea.
Organizational Affiliation: