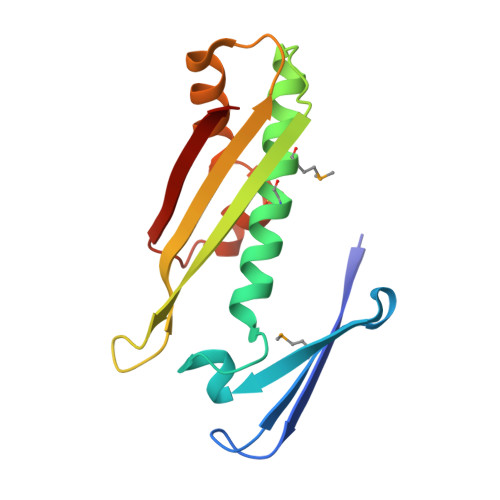
Crystallographic Structure and Biochemical Analysis of the Thermus thermophilus Osmotically Inducible Protein C
Rehse, P.H., Oshima, N., Nodaka, Y., Tahirov, T.H.(2004) J Mol Biology 338: 959-968
- PubMed: 15111059 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.03.050
- Primary Citation Related Structures:
1UKK - PubMed Abstract:
The X-ray crystallographic structure of osmotically inducible Protein C from the thermophilic bacterium, Thermus thermophilus HB8, was solved to 1.6A using the multiple wavelength anomalous dispersion method and a selenomethionine incorporated protein (Se-MAD). The crystal space group was P1 with cell dimensions of a=37.58 A, b=40.95 A, c=48.14 A, alpha=76.9 degrees, beta=74.0 degrees and gamma=64.1 degrees. The two tightly interacting monomers in the asymmetric unit are related by a non-crystallographic 2-fold. The dimer structure is defined primarily by two very long anti-parallel, over-lapping alpha-helices at the core, with a further six-stranded anti-parallel beta-sheet on the outside of the structure. With respect to the beta-sheets, both A and B monomers contribute three strands each resulting in an intertwining of the structure. The active site consists of two cysteine residues from one monomer and an arginine and glutamic acid from the other. Enzymatic assays have revealed that T.thermophilus OsmC has a hydroperoxide peroxidase activity.
- Highthroughput Factory, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: