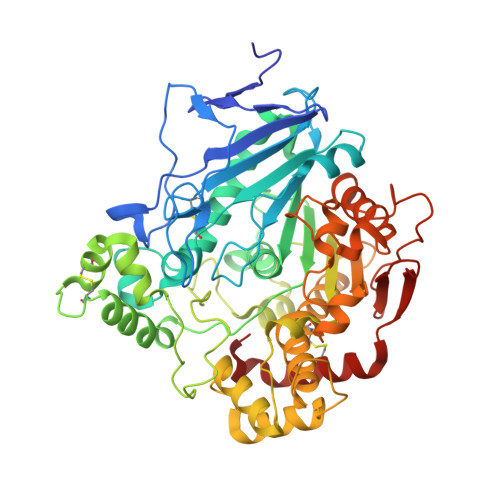
Aspergillus niger protein EstA defines a new class of fungal esterases within the alpha/beta hydrolase fold superfamily of proteins
Bourne, Y., Hasper, A.A., Chahinian, H., Juin, M., De Graaff, L.H., Marchot, P.(2004) Structure 12: 677-687
- PubMed: 15062090 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.03.005
- Primary Citation Related Structures:
1UKC - PubMed Abstract:
From the fungus Aspergillus niger, we identified a new gene encoding protein EstA, a member of the alpha/beta-hydrolase fold superfamily but of unknown substrate specificity. EstA was overexpressed and its crystal structure was solved by molecular replacement using a lipase-acetylcholinesterase chimera template. The 2.1 A resolution structure of EstA reveals a canonical Ser/Glu/His catalytic triad located in a small pocket at the bottom of a large solvent-accessible, bowl-shaped cavity. Potential substrates selected by manual docking procedures were assayed for EstA activity. Consistent with the pocket geometry, preference for hydrolysis of short acyl/propyl chain substrates was found. Identification of close homologs from the genome of other fungi, of which some are broad host-range pathogens, defines EstA as the first member of a novel class of fungal esterases within the superfamily. Hence the structure of EstA constitutes a lead template in the design of new antifungal agents directed toward its pathogenic homologs.
- Architecture et Fonction des Macromolécules Biologiques, CNRS UMR-6098, 31 Chemin Joseph Aiguier, F-13402 Marseille Cedex 20, France. yves@afmb.cnrs-mrs.fr
Organizational Affiliation: