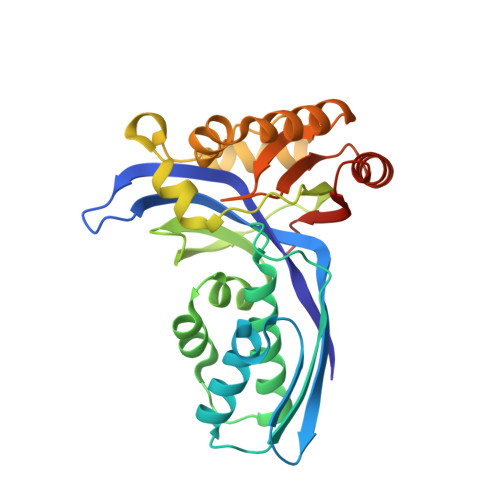
Crystal Structure of 4-(Cytidine 5'-diphospho)-2-C-methyl-D-erythritol kinase, an Enzyme in the Non-mevalonate Pathway of Isoprenoid Synthesis.
Wada, T., Kuzuyama, T., Satoh, S., Kuramitsu, S., Yokoyama, S., Unzai, S., Tame, J.R., Park, S.Y.(2003) J Biological Chem 278: 30022-30027
- PubMed: 12771135 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304339200
- Primary Citation Related Structures:
1UEK - PubMed Abstract:
The crystal structure of the enzyme 4-(cytidine 5'-diphospho)-2-C-methyl-D-erythritol (CDP-ME) kinase from the thermophilic bacterium Thermus thermophilus HB8 has been determined at 1.7-A resolution. This enzyme catalyzes phosphorylation of the 2-hydroxyl group of CDP-ME, the fourth step of the non-mevalonate pathway, which is essential for isoprenoid biosynthesis in several pathogenic microorganisms. Since this pathway is absent in humans, it is an important target for the development of novel antimicrobial compounds. The structure of the enzyme is similar to the structures of mevalonate kinase and homoserine kinase, members of the GHMP superfamily. Lys8 and Asp125 are active site residues in mevalonate kinase that also appear to play a catalytic role in CDP-ME kinase. Both the mevalonate and the non-mevalonate pathways therefore involve closely related kinases with similar mechanisms. Assaying the enzyme showed that CDP-ME kinase will phosphorylate CDP-ME but not 4-(uridine 5'-diphospho)-2-C-methyl-D-erythritol, indicating the substrate pyrimidine moiety is involved in important interactions with the enzyme.
- Genomic Sciences Center, RIKEN Yokohama Institute, and Protein Design Laboratory, Yokohama City University, 1-7-29 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: