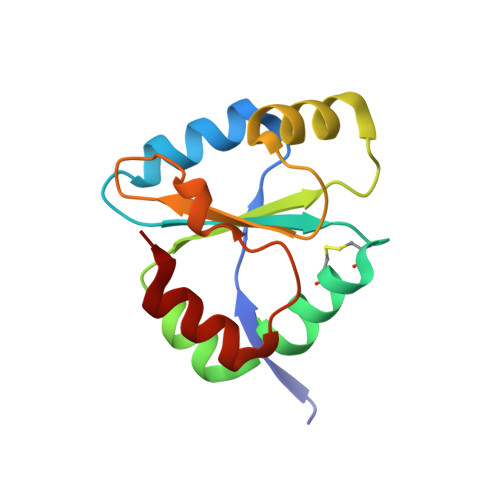
Crystal structure of DsbDgamma reveals the mechanism of redox potential shift and substrate specificity(1)
Kim, J.H., Kim, S.J., Jeong, D.G., Son, J.H., Ryu, S.E.(2003) FEBS Lett 543: 164-169
- PubMed: 12753926 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(03)00434-4
- Primary Citation Related Structures:
1UC7 - PubMed Abstract:
The Escherichia coli transmembrane protein DsbD transfers electrons from the cytoplasm to the periplasm through a cascade of thiol-disulfide exchange reactions. In this process, the C-terminal periplasmic domain of DsbD (DsbDgamma) shuttles the reducing potential from the membrane domain (DsbDbeta) to the N-terminal periplasmic domain (DsbDalpha). The crystal structure of DsbDgamma determined at 1.9 A resolution reveals that the domain has a thioredoxin fold with an extended N-terminal stretch. In comparison to thioredoxin, the DsbDgamma structure exhibits the stabilized active site conformation and the extended active site alpha2 helix that explain the domain's substrate specificity and the redox potential shift, respectively. The hypothetical model of the DsbDgamma:DsbDalpha complex based on the DsbDgamma structure and previous structural studies indicates that the conserved hydrophobic residue in the C-X-X-C motif of DsbDgamma may be important in the specific recognition of DsbDalpha.
- Center for Cellular Switch Protein Structure, Korea Research Institute of Bioscience and Biotechnology, 52 Euh-eun-dong, Yusong-gu, 305-806, Daejon, South Korea.
Organizational Affiliation: